Abstract
OBJECTIVE To investigate the expression of the RECK gene in human breast (cancer) cell lines, and to determine the relationship between RECK gene expression and the invasive capacity of the breast cancer cell lines.
METHODS The invasive capacity of breast (cancer) cell lines including HBL-100, MCF-7 and MDA-MB-435S were determined by the Transwell method. The protein expression levels of RECK, MMP-2 and MMP-9 genes in these three cell lines were measured by immunocytochemical methods. The expressions of the RECK gene and protein level were measured by RT-PCR and Western blots in the cell lines respectively.
RESULTS The order of the invasive capacity of the breast (cancer) cell lines was MDA-MB-435S, being the highest, and HBL-100, being the lowest. The invasive capacity difference between any two groups among the three groups was significant (P<0.01). The protein expression level of the RECK gene in the HBL-100 cell line was highest, and no expression was detected in MDA-MB-435S cells. Moreover, the expression of the RECK gene was negatively correlated with the expression of the MMP-2 and MMP-9 genes. The mRNA level of the RECK gene in HBL-100 cells was the highest, but no expression was found in the MDA-MB-435S cells (P<0.001).
CONCLUSION There was a significant negative correlation between the expression level of the RECK gene and invasive capacity in vitro, and the RECK gene expression showed an inverse proportion to that of the MMP-2, MMP-9 genes.
keywords
The degradation of the extra-cellular matrix plays a key role in the process of cancer growth, parenchymal invasion, metastasis and angiogenesis. Cancer cells degrade the extra-cellular matrix with the help of MMP (matrix metallproteinase) to expedite their infiltration and metastasis. Research shows there is a relationship between breast cancer progression and the increase of MMP expression.[1,2] Inhibitors of MMP are regarded as promising agents to inhibit cancer growth and metastasis as a result of their inhibitory activity on angiogenesis.[3] One of the recently found inhibitors is RECK (reversion-inducing-cysteine-rich protein with Kazal motifs), thought to be a newly discovered cancer prognosis gene. In cases of pancreatic carcinoma, those with a high expression of RECK have a better prognosis than those with lower expression.[4] However in cases of breast cancer, this relationship between the RECK and MMP genes expression, relative to invasive capacity was not found. Therefore, we attempted to examine the correlation between the invasive capacity and the expressions of the RECK gene in three breast (cancer) cell lines with different inva-sive capacities.
MATERIALS AND METHODS
Cell lines and cell culture
HBL-100 cells are human breast non-tumor cells; MCF-7 cells are human breast adenocarcinoma cells and MDA-MB-435S cells are human breast ductal carcinoma cells. All cells were maintained in 25-cm2 flasks in a humidified 5% C02/95% air incubator at 37°C in RPMI 1640 medium (Life Technologies, Inc.) supplemented with 2 mM glutamine, 100 units/ml penicillin, 100 ug/ml streptomycin, and 10% heat-inactivated fetal bovine serum (FBS, Hyclone laboratories, Inc., Logan, UT). Cell counts were performed with the aid of a hemocytometer in the presence of 0.02% trypan blue to assess cell viability.
Cell invasion assay
Transwells (Coming Costar) with 12-µm polycarbonate filters, 12-μm pore size, were used. Twenty-five microliters of 100 (xg/ml fibronectin (Sigma) was applied on the lower side of the filters, which were left for 1 h in a laminar hood to dry. The upper side of the filters was coated with 100 µl of 1 mg/ml Matrigel (Becton Dickinson) and 100 µl of RPMI 1640 medium was added. The Matrigel was dried overnight at room temperature in a laminar hood and reconstituted with 200 JJLI of mdium for 1 h at 37°C. The upper compartments were filled with 200 pi of cell suspension, final concentration lxl05/ml, and the lower compartments were filled with 800 pi medium. The plates were incubated for 24 h at 37°C in 5% C02. The cells on the upper side of the filters were scraped and the wells put upside down. The filters were fixed using methanol and stained with Giemsa. The number of cells invaded into the filters was counted. All tests were performed in triplicate.
Immunocytochemical procedures
The cells were seeded on glass coverlips and allowed to grow until half confluence. They were washed with PBS three times and then fixed by immersion in cold acetone for 10 min. After several washes in PBS, the cells were treated with 0.3% hydrogen peroxide for 10 min. After rinsing, the cells were blocked with non-immune animal serum, 50 μ1 for 15 min and subsequently incubated overnight at 4°C in a moist chamber with primary antibodies (anti-RECK, from BD Biosciences Inc., USA; anti-MMP-2 and anti-MMP-9, from Zhongshan Bio-technique Inc., Beijing, China) diluted in PBS. After three washes with PBS, the slides were incubated for 1 h at 37°C with secondary antibodies (rabbit anti-mouse IgG, Zhongshan Bio-technique Inc.), diluted and washed in PBS and incubated with the avidin-biotin-peroxide reagents (Zhongshan Bio-technique Inc.) for 30 min at 37°C. Then the slides were washed in PBS, and the antigen-antibody complex visualized using diaminobenzidine (Stable DAB, Zhongshan Bio-technique Inc.). Negative controls were treated identically with omission of the primary antibody. The immunocytochemical reactions were visualized using Olympus AX70 equipped with a 400 x magnification. Cells were scored as positive for membrane or cytosol immunoperoidase stain by two independent observers. The RECK, MMP-2 and MMP-9 protein expression levels in these three cell lines were calculated using an image analysis program, Metamorph 4.5 (Universal Imaging Corp., West Chester, PA) by taking two slides from each cell line, and measuring the OD values of the 5 same areas on each slide. The protein level was proportional to the OD value.
Western blot analysis
Initially, the cells were washed twice in ice-cold PBS. After this step, total cell extracts were prepared from cells which were lysed in 50 µl of RIPA buffer (50 mM Tris, pH 8.0, 100 mM NaCl, 0.1% (w/v) SDS, 1% (v/v) Nonidet P-40, 0.5% (w/v) deoxycholic acid and 1 mM EDTA). Following 10 min of incubation on ice, insoluble material was removed by centrifugation at 14,000 rpm for 10 min. The supernatants were transferred to fresh test tubes and the total protein concentration was determined (Bradford assay Bio-Rad). The protein (50 µg) of whole cell lysates was resolved by 8% SDS-PAGE, and subsequently transferred to polyvinylidene difluoride membranes (Millipore, Bedford, MA) in a tranfer buffer (39 mM glycine, 48 mM Tris, 0.037% SDS, and 15% methanol). These membranes were blocked in PBS containing 5% non-fat milk for 1 h. They were then washed in PBS containing 0.1% Tween 20 (Bio-Rad) three times. Mouse anti-RECK antibody (1:250) and goat anti-actin(l :500) were used as primary antibodies diluted in 0.1% PBST (phosphate buffered saline containing 0.1% Tween 20) overnight at 4°C. After that step, they were washed in 0.1% PBST several times. Rabbit anti-mouse and rabbit anti-goat IgG antibodies were used as secondary antibodies at a 1:5000 dilution for 30 min. Chemiluminescence (Amersham Biosciences) was then used to visualize immunoreactive protein complexes. The exposure time for the detection of RECK and actin was 1min. Densitometrie scanning of the bands was used for quantitation using a gel scanning image analysis system, Chemimage 5500 (Alpha Inno Tech.), and data were normalized to ß-actin expression as a RECK protein mRNA index.
Semi-quantitative RT-PCR analysis
Total RNA was isolated from these three cell lines by the guanidium thiocyanate method using a Trizol reagent (Gibco-BRL). Total RNA was reverse transcribed into cDNA (Bao Biotech Inc.) in a 20μl final volume, followed by PCR. The primers for RECK and ß-actin were designed on the basis of established Genbank sequences. The effective primers for RECK were as follows: 5’-AAG ACA GCT CGC GTG TGA CC (sense) and 5'-TGG GAT GAT GGG TTT GCA TC (antisense). The expected DNA length was 348 bp. ß-actin: 5'-GTG GGG CGC CCC AGG CAC CA (sense) and 5'-CTC CTT AAT GTC ACG CAC GAT TTC (antisense). The expected DNA length was 498 bp. Conditions for DNA amplification included an initial dénaturation step of 5 min at 94°C and 30 cycles of 1 min at 94°C 30 s at 58°C and 40 s at 72°C and extension for 10 min at 72°C. PCR samples were electrophoresed in a 1.5% agarose gel and stained with ethidium bromide (0.5 (xg/ml). Densitometrie scanning of the bands was used for quantitation using a gelscanning image-analysis system, Chemimage 5500, and data were normalized to ß-actin expression as a RECK mRNA index.
Statistical analysis
All statistical analyses were performed using the SPSS 10.0 statistical package. Data were expressed as the mean±SEM. One-way ANOVA analysis was used to compare differences among the means.
RESULTS
The invasive capacities of breast non-cancer and cancer cells
Invasive capacity in vitro is measured by the number of cells penetrating the filter membranes. The order of the invasive capacity of breast (cancer) cell lines was: MDA-MB-435S, being the highest (425.20 ±54.09); MCF-7 (239.00±91.39) and HBL-100 (101.00±63.88), the lowest. The invasive capacity differences between any two groups among the three groups were significant (P<0.01).
The level of the RECK gene protein expression in the three groups
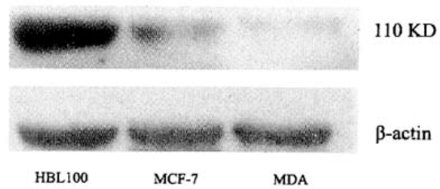
The level of the RECK gene protein expression in HBL-100 cells was highest (IC: 3188.24±894.86, WB: 3.32±0.25), in MCF-7 was (IC: 1058.92±336.53, P< 0.01, WB: 2.23±0.59, P<0.05), and no expression was found in the MDA-MB-435S cell line. Moreover, the expression of the RECK gene was negatively correlated with the expression of MMP-2 and MMP-9 genes (Table 1 and Fig. 1).
RECK expression in breast (cancer) lines.
The level of RECK gene mRNA expression in the three groups
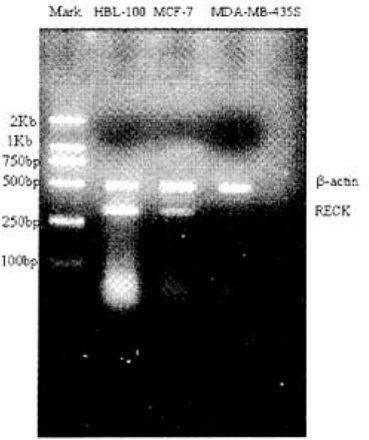
RT-PCR was used to measure the level of RECK gene mRNA expression in the three groups. Table 1 and Fig. 2 show that the mRNA level of the RECK gene in HBL-100 cells was the highest (2.32±0.02) with no expression in MDA-MB-435S cells (P<0.001).
RECK mRNA level in breast (cancer) cell lines.
DISCUSSION
The RECK gene is a transformation suppressor gene newly discovered in recent years, and its presence has been verified by a screen of a human fibroblast cDNA expression libraries. This gene can induce flat morphology (or "flat reversion") when expressed in a v-Kiras-transformed NIH 3T3 cell line. RECK is a membrane-anchored glycoprotein of 110 kDa in size, located at 9pl3-pl2. It is longer than 87 Kb,[5] contains many epidermic growth factor-like repeats and serine protease-inhibitor-like domains, but has no structural homology with TIMPs. RECK mRNA is widely expressed in various normal human tissues and untransformed cells while it is undetectable in tumor-derived cell lines and oncogenically-transformed cells.[6]
The invasion of tumor cells into the basement membrane is the key stage of metastasis and directly affects the prognosis. The test of invasive capacity of tumor cell penetration into an artificial basement membrane is significant, as this assay indirectly reflects its invasive capacity in vivo. The three groups of breast (cancer) cells represent three types, of which invasive capacities are verified to be different through the TRANSWELL method. MDA-MB-435S cells showed the highest invasiveness, while HBL-100 cells displayed the lowest. Furthermore, the immunocytochemical technique, Western blot and RT-PCR methods were used to test the RECK gene protein and the expression level of mRNA respectively for the three cell lines. The results show that the highly invasive MDA-MB-435S cells had no expression of the RECK gene while the lowest invasive HBL-100 cells had the highest expression levels of both the RECK mRNA and proteins. Therefore, the expression of the RECK gene can inhibit the invasive capacity of breast cancer cells in vitro, as they are negatively correlated. The function of the RECK gene to inhibit the cancer cell invasive capacity is probably related to its inhibitory effect on the secretion and activity of MMP-2, MMP-9 and MT1-MMP and thereby suppresses tumor angiogenesis.[5,7] The inhibitive role of RECK expression against the protein expression of MMP-2 and MMP-9 was also verified in this study.
Animals with a higher expression of RECK genes have been shown to live longer than the ones in the control group without the expression of that gene.[7] In addition, among various hepatic cancer cells, the expression level of the RECK gene is apparently correlated with time of patient survival.8 Therefore, RECK gene expression can be regarded as an indicator for prognosis of cancer sufferers. Because expression of the RECK gene expression in cancer tissues affects growth, invasion, metastasis and angiogenesis, enhancement of its expression may be expected to be a new target for tumor prevention and therapy.
- Received July 17, 2006.
- Accepted September 26, 2006.
- Copyright © 2006 by Tianjin Medical University Cancer Institute & Hospital and Springer