Abstract
OBJECTIVE Telomerase is an attractive molecular target for cancer therapy because the activation of telomerase is one of the key steps in cell immortalization and carcinogenesis. RNA Interference using small-Interfering RNA (siRNA) has been demonstrated to be an effective method for Inhibiting the expression of a given gene in human cells. The aim of the present study was to investigate whether inhibition of telomerase activity by siRNA targeted against human telomerase RNA (hTR) can inhibit proliferation and induce apoptotic cell death In human renal carcinoma cells (HRCCs).
METHODS The siRNA duplexes for hTR were synthesized and 786-0 HRCCs were transfected with different concentrations of hTR-siRNA. The influence on the hTR mRNA level, telomerase activity, as well as the effect on cell proliferation and apoptosis was examined.
RESULTS Anti-hTR siRNA treatment of HRCCs resulted In specific reduction of hTR mRNA and inhibition of telomerase activity. Additionally, significant Inhibition of proliferation and Induction of apoptosis were observed.
CONCLUSION siRNA against the hTR gene can Inhibit proliferation and induce apoptosis by blocking telomerase activity of HRCCs. Specific hTR inhibition by siRNA represents a promising new option for renal cancer treatment.
keywords
The incidence of renal cell carcinoma (RCC), the most common malignant disease of the adult kidney, has steadily increased. A lack of effective treatment for RCC is largely due to the resistance of the cancer cells to conventional modes of treatment, such as radiotherapy and chemotherapy.[1] Therefore, new therapeutic approaches are required for RCC.
It has been shown that the introduction of double-stranded oligori-bonucleotides, also called small-interfering RNA (siRNA) into mammalian cells, triggers the degradation of the endogenous mRNA to which the siRNA hybridizes.[2] Many studies [3,4] have demonstrated that the transfer of siRNA into cells is an effective method for inhibiting the expression of oncogenes. siRNA cell transfer is currently being evaluated as a potentially useful method for genetic-based cancer therapy.
Human telomerase is a specialized reverse transcriptase that catalyses telomeric repeat addition at the ends of chromosomes.[5] Telomerase consists of a protein component, hTERT, and an RNA component (hTR) containing the template for adding repeating units onto the ends of chromosomes.[6] The ability of cells to replicate indefinitely has been linked to telomerase expression.
A high percentage of tumor cells that take on immortalized characteristics show telomerase activity. Hara et al.[7] found that elevated levels of telomerase activity correlated with tumor stages as well as with the degree of nuclear RCC grades. The ubiquitous expression of telomerase in human tumors, including RCC, has supported the hypothesis that the enzyme is involved in cellular immortality and carcinogenesis. It has been demonstrated that activation of telomerase is one of the key steps in cell immortalization and carcinogenesis.[8]
In the present study, using a human renal carcinoma cell line, we evaluated the inhibitory effect of siRNA directed against the hTR region containing the telomere repeat template sequence on telomerase activity. Incubation of the human 786-0 RCC with appropriate siRNAs resulted in inhibition of hTR mRNA expression and telomerase activity. siRNA-mediated inhibition of telomerase activity subsequently led to significiant inhibition of proliferation and apoptotic cell death.
Materials and Methods
siRNA preparation
The siRNAs duplexes were synthesized, purified, and annealed by the Ambion Co. (USA). The siRNA targeted the hTR region containing the telomere repeat template sequence: sense sequence 5‘ -UUG UCU AAC CCU AAC UGA GTT-3 ’ and antisense sequence 3‘ -TTA ACA GAU UGG GAU UGA CUC-5’. The selected sequence was submitted to BLAST search to assure that the only, the selected gene was targeted. A scrambled siRNA was purchased from Ambition (silencer™ Control siRNA#3) and used as a control.
Cell culture and transfection
Human renal carcinoma 786-0 cells were obtained from the Shanghai Cell Institute (Shanghai, China) and cultured in RPMI-1640 medium supplemented with 10% FCS, penicillin and streptomycin. The cells were routinely passaged to maintain exponential growth. The day before transfection, the cells were trypsinized, diluted with fresh medium and transfered to 24-well plates. Transfection of siRNAs was carried out using siPORT™ lipid (Ambion). siPORT lipids and siRNAs were both diluted into OPTI-MEM I, respectively. Diluted siPORT lipids were mixed with diluted siRNAs and the mixture was incubated for 20 min at room temperature for complex formation. After addition of OPTI-MEM I to each well containing cells to a level of 200 pi, the entire mixture was added to the cells in one well resulting in a final concentrations of 10, 50 and 100 nM siRNAs. Cells were harvested and assayed at 24, 48 and 72 h after transfection. All experiments were repeated at least six times.
RT-PCR
Analysis of hTR RNA was performed by RT-PCR amplification. Total RNA was purified using a Total RNA isolation system and RT-PCR was performed using the Acess RT-PCR system (Promega). The upstream and downstream primers were 5‘-CTG GGA GGG GTG TG G CCA TTT-3’ and 5‘-CGA ACG GGC CAG CAG CTG ACAT-3’, respectively. Reaction parameters were 94°C for 20 s, 50°C for 20 s, and 72°C for 30 s for 25 cycles. GAPDH was used as an internal control to assure the accuracy. Quantitation was performed with an image analyzer (LabWorks Software, UVP Upland, CA, USA).
In situ hybridization
Chamber slides were permeabilized with proteinase K digestion (2.5 μg/ml) followed by acetylation. Hybridization was performed overnight at 37°C with a human hTR-biotinylated DNA probe that was a 30-mer oligonucleotide (5‘-TTC TAC CGG AAG AGT TCG TGA GCA AGT TG-3’), which was synthesized and biotinylated by the Boshide Biotechnology Co (Wuhan China). After post-hybridization washes, the signals were detected immunochemically by subsequent incubation with a streptavidin-horseradish peroxidase conjugate and developed by the DAB system.
Telomerase activity
Telomerase activity assay was performed according to a polymerase chain reaction-based telomeric repeat-amplification protocol (TRAP) as described by Mehle [9] using the Telomerase PCR ELISA Kit (Roche, ermany) following the provider's instructions.
TUNEL assay
The TUNEL technique was performed to detect and quantify apoptotic cell death using the In situ Cell Death Detection Kit (Roche Diagnostics, USA) following the instructions supplied by the manufacturer. Briefly, chamber slides were fixed with 4% paraformaldehyde for 30 min and permeabilized in 0.1% Triton-100, 0.1% sodium citrate at 4°Cfor 2 min. The slides were incubated with the TUNEL reaction mixture for 1 h at 37°C. After washing with PBS, the slides were incubated with peroxidase-conjugated antibody for 30 min at 37°C and were developed with the DAB system. Microscopically, six fields were randomly selected from every sample, then 100 cells were randomly selected from every field. The apoptotic rate= (number of total apoptotic cells/100) × l00%.
Cellular growth curve
For evaluation of cell numbers, the cells were trypsinized at the appropriate times, stained with trypan blue and counted using a hemocytometer. Each experimental condition was performed six times, and the average value for each group was determined to compose the growth curve.
Cell proliferation assay
Cell proliferation was assayed by the MTT method. In brief, the 786-0 cells (2 × l04 cells/well) were incubated in a 96-well plate, in the absence or presence of siR-NA, at 37°C in a humidified atmosphere containing 5% CO2. At the end of the experiment, 20 μ1 of MTT dye (3-4,5-dimethylthiazol zyl-diphenyltetrazolium bromide, 5 mg/ml) (Sigma, USA) was added to each well. Four hours later, 100 μ1 of DMSO per dish was added and absorption units (UA) determined at 570nm on a ELX-800 spectrometer reader (Bio-Tek instruments Inc, USA).
Proliferation inhibition rates (%)= (l-UAE/UAC) × 100%.
UAE: average UA value of experimental group;
UAC: average UA value of control group.
Statistical analysis
Values were expressed as the mean±SD and obtained from at least six independent groups (n=6). Statistical analysis of the results was carried out by one-way analysis of variance (ANOVA) followed by the Duncan's new multiple range method or Newman-Keuls test. PÀ.05 were considered significant.
RESULTS
Effect of siRNA treatment on hTR mRNA expression
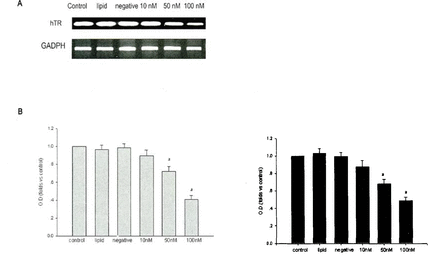
hTR mRNA expression was examined using RT-PCR and in situ hybridization at 24 h. As shown in Fig.l, the 786-0 cells treated with hTR siRNAs (50, 100 nM) displayed significantly decreased hTR mRNA levels compared to 786-0 cells treated with negative control siRNAs in the RT-PCR assay (Fig.lA, B) and by in situ hybridization (Fig. 1C)
Effect of siRNA on hTR mRNA expression.The 786-0 cells were transfected with siRNAs at the concentrations shown. After 24 h, the cells were harvested and hTR mRNA levels were quantified by RT-PCR (A) and by in situ hybridization. Transfection with hTR siRNA significantly decreased the amount of hTR mRNA detectable in the 786-0 cells. Quantified analysis by RT-PCR (B) and in situ hybridization(C). Data are expressed as the mean± SD and as the folds name vs. the control. aP<0.01 vs. control (n=6).
Effect of siRNA treatment on telomerose activity
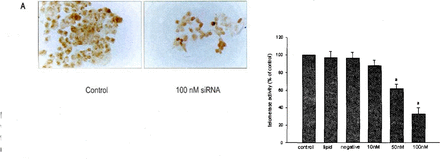
The efrect of the hTR siRNAs on telomerase activity was performed by the TRAP assay at 48 h.Results throughout are reported as a percentage of telomerase activity of untreated cells(contro1).As shown in Fig.2. siRNAs for hTR depressed the telomerase activity of the 786.0 cells in a concentration.dependent manner. Cells treated with hTR siRNAs(50.100 nM)exhibited significantly decreased telomerase activity compared to the 786.0 cells treated with negative-contro1 siRNAs.The maximum effect observed with 1 00 nM hTR siRNA reduced the activity to 33%of the untreated cells(Fig.2).
Effect of siRNA on telomerase activity.The 786-0 cells were transfected with siRNA al concentrations shown.After 48 h,the cells were harvested and telomerase activity was quantified by the TRAP assay.Results throughout are reposed as a percentage of telomerase activity of untreated cells(contro1).
Apoptotic cell death
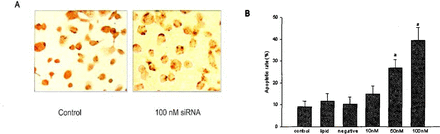
When apoptosis of the 786-0 cells was evaluated by the TUNEL technique, about 10% of cells cultured with negative-control siRNAs manifested evidence of apoptotic change after 72 h in culture. In contrast, a significantly greater proportion (approximately 39%) of the 786-0 cells cultured with 100 nM hTR siRNA were TUNEL positive (Fig.3).
TUNEL staining showing apoptotic changes in the 786-0 cells. (A) Only occasional cells transfected with control siRNA show positive TUNEL staining. In contrast, the majority of cells transfected with hTR siRNA demonstrate positive TUNEL staining. (B) Group comparisons of TUNEL assay illustrating the effects of transfection with hTR siRNA.
Antiproliferative effects of hTR siRNAs treatment
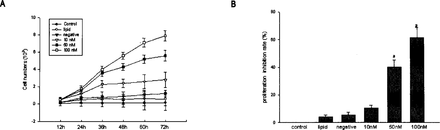
Cell numbers were determined on days 1~3 following siRNAs treatment. hTR siRNA (50, 100 nM) treatment resulted in a marked inhibition of cellular proliferation over this 3-day period. Cell growth was not influenced significantly by treatment with control siRNAs or with the concentration-transfection regent control. The siR-NA-induced anti-proliferative effect was dose-dependent (Fig.4A). In the MTT proliferation assay of the 786-0 cells, proliferation, treatment with hTR siRNAs (100 nM) caused a decreased of 63% compared to the control group (Fig.4B).
Effect of siRNA on 786-0 cell growth.(A)The 786-0 cells were treated with lipid,negative siRNA (100 nM)and siRNA (10,50 and 100 nM)and then trypsinized and counted by a hemocytometer at the appropriate time. (B)Transfection with hTR siRNA significantly increased the proliferation inhibition rates of 786-0 cells determined by the MTT assay.
Discussion
Currently there are no effective treatment options for RCC because it is relatively resistant to radiotherapy and chemotherapy, and responses to immunotherapy are poor.[1] In the search for a novel therapeutic approach, the inhibition of telomerase activity in RCC cells has gained considerable interest.
The main function of telomerase is to catalyze lengthening and capping of the ends of telomeres.[5] Uncapped or critically shortened telomeres cause inhibitory cellular responses, such as cell cycle arrest and apoptosis, 1101 and depressed telomerase limits growth of human cancer cells.[11] Moreover, more than 90% of examined tumors including RCCs show telomerase activity.1121 Mehle et al.[9] tested 56 RCCs for telomerase activity and found 40 of the analyzed tumors (71%) were positive, whereas none of 56 corresponding normal kidney samples showed telomerase activity.
A strategy to inhibit telomerase activity by phospho-rothioate-modified oligonucleotids has been reported. 1131 However, the poor sequence selectivity observed with such compounds has lead to the application of second-generation oligonucleotides, peptide nucleic acids (PNAs). One study showed that PNAs targeting human telomerase inhibited telomerase activity at a level that was 10-50 fold more efficient than inhibition by analogous oligonucleotids.[14] While solid evidence of antisense effects of PNAs has been demonstrated, using PNAs as gene-therapeutic drugs has been hampered by poor intrinsic uptake of PNAs by living cells. [15] Recently, the demonstration that RNA interference (RNAi) can be used to inhibit gene expression in mammals opens new avenues for gene-targeted therapies.[16]
RNAi is a sequence-specific, post-transcriptional gene silencing mechanism. By the introduction of small-interfering RNA (siRNA), a hybrid is formed consisting of a sense and antisense strand homologous in sequence to the silenced gene.[2] siRNA, 21-ntRNA, with 2-nt 32 overhang can mediate strong and specific suppression of gene expression.[17] Takei et al.[18] reported that the siRNA targeting human VEGF almost com pletely inhibited the secretion of VGEF in the human prostate PC3 cancer cell line. The VEGF siRNA dramatically suppressed tumor antiogenesis and tumor growth in a PC3-xenograft model.
In the present study, we evaluated the ability of siRNA targeting hTR to reduce the telomerase activity. When the human renal carcinoma 786-0 cells were transfected with hTR siRNA, the hTR mRNA level of the 786-0 cells decreased. siRNA inhibited hTR expression in a concentration-dependent fashion. Moreover, the specificity of the siRNA for the hTR target gene was demonstrated by the inability of control siRNA to inhibit hTR expression. Our results also showed that telomerase activity in human cancer cells can be inhibited by hTR siRNA in a concentration-dependent manner. The maximum effect observed in the 786-0 cells was 33% of untreated cell telomerase activity. A reduction of the hTR mRNA amount and telomerase activity after anti-hTR treatment also was observed by Kosciolek et al.,[19] in HCT-15 human colon carcinoma cells and in HeLa cervical carcinoma cells. However, it should be emphasized that in the 786-0 cells, hTR expression and telomerase activity were markedly reduced but not completely eliminated. The exact site targeted by the RNA may be a key factor because some sites are bonded internally or to neighboring molecules.[20]
Our study has shown that the reduction of hTR expression and telomerase activity was closely associated with or rather followed by apoptotic changes. The 786-0 cells transfected with a high concentration of hTR siRNA exhibited lower hTR expression and telomerase activity and a higher degree of apoptosis. A low concentration of hTR siRNA had only a minor effect on hTR expression and telomerase activity and apoptotic induction. We also have shown that application of hTR siRNA depresses cell proliferation. The siRNA-induced anti-proliferative effects are dependent on the reduction of hTR expression and reduction of telomerase activity.
It is thought that the time necessary to observe decreased proliferation after siRNA treatment should vary depending on initial telomere length. A celluar response that depends on telomere shortening will require weeks to become apparent.[21] But other studies have shown immediate cellular responses to telomerase inhibition independent of telomere length.[22,23] These immediate effects appear to be caused by deficient telomere capping. We propose that the immediate effects on cell growth inhibition shown in our study are based on the deficient telomere capping mechanism. A rapid response to telomerase inhibition on growth of ovarian cancer cells was shown by Saretzki et al.[22] applying ribozymes, and rapid responses of telomerase inhibition on apoptosis of bladder cancer cells were shown by Kraemer et al.[20] applying antisense oligonucleotids. The loss of telomere integrity and, as a consequence, the accumulation of DNA damage resulted in cell-cycle arrest at G1 and apoptosis.[22]
Overall, our results suggest that hTR plays an essential role in cell proliferation and viability control of human renal carcinoma cells. RNAi represents a new and powerful gene-silencing approach that is currently believed to be more efficacious, selective, and specific than antisense techologies.
Acknowledgements
We thank Zhang GY for very useful comments and Hu SQ for technical assistance.
Footnotes
This work was supported by grants from the Jiangsu Science and Technology Department (No.BK2005429)and Health Departmental Science Research Funds of China (No.2005-05).
- Received July 5, 2006.
- Accepted October 10, 2006.
- Copyright © 2006 by Tianjin Medical University Cancer Institute & Hospital and Springer