Abstract
OBJECTIVE Because almost all malignancies represent monoclonal proliferations, we have studied the clonal status of peripheral papillomas (peri-PM), ductal carcinomas in situ (DCIS), and normal breast tissues to explore a reliable way to distinguish benign and malignant (or pre-malignant) cases previously diagnosed morphologically.
METHODS Twenty-six cases of peri-PM, 25 cases of peri-PM with atypical ductal hyperplasia (ADH), 27 cases of DCIS, 16 cases of developed canceration and 20 specimens of normal tissue were examined in the study. The clonal status of these tissues was studied using an assay based on inactivation mosaicism of the lenth–polymorphic X–chromosomes at the androgen receptor (AR) locus.
RESULTS Loss of polymorphism at the AR locus was found in all DCIS cases and 10 cases (10/25, 40.0%) of peri-PM with ADH, indicating the monoclonality of the tumors. Twenty-four out of 26 (92.3%) cases with peri-PM and 19 specimens of normal tissue were shown to be polyclonal. In 16 cases of developed canceration, identical X chromosome inactivation (monoclonal alterations) was observed from both the peri-PM with ADH part, and the DCIS part in each case.
CONCLUSION These results contribute to the understanding of the genetic changes of peri-PM, and confirm the peri-PM with ADH as a precancerous lesion of the breast. Clonal analysis might be a useful modality to screen high-risk cases from precancerous lesions or to distinguish between benign hyperplasia and early carcinoma.
keywords
- peripheral papilloma of the breast
- precancerous lesion
- ductal carcinoma in situ
- clonal analysis
- androgen receptor
INTRODUCTION
The existing system for the classification of intraductal epithelial proliferations assumes a spectrum from usual epithelial hyperplasia through ADH and low-grade DCIS to high-grade disease[1]. It is now generally agreed that the risk of breast carcinoma development is estimated to be 1.5 times that of the reference population for usual ductal hyperplasia, 4~5 fold for ADH and 8~10 fold for DCIS[2]. But a definition of a premalignant morphologic continuum is lacking. Therefore, we attempted to provide a classification system for epithelial proliferating lesions and in situ malignancy of the breast. Cancer and premalignant lesions have been demonstrated to share several genetic alterations[3-6]. It would be desirable to develop a molecular assay that could detect biologic alterations that would allow definitive diagnosis of malignancy, even without morphologic inspection.
Many new studies have apparently confirmed that the monoclonal origin from the monocytohyperplasia in human epithelial (and other) tumors is an important feature of malignant tumors. It has been indicated by some reports that in normal epithelial cells and benign proliferated lesions, proliferation is usually polyclonal; Ductal carcinomas almost always display monoclonal proliferation[7-10]. However, in Chinese women it is not clear to the clonality of breast peripheral papillomas (peri-PM)[11], peri-PM with atypical ductal hyperplasia (ADH), developed cancerous cells and ductal carcinoma in situ (DCIS). So, in our study clonal analyses were performed with 4 groups of small DNA samples from paraffin-embedded tissue of peri-PM, peri-PM with ADH, developed cancerous cells, and DCIS. Twenty normal breast tissues were used as a control group.
The method used for clonal analysis[8,12] was based on restriction fragment length polymorphism of the female X-chromosome-linked androgen receptor (AR) gene, and on the differential methylation of the AR gene due to random inactivation of one of two X-chromosomes by methylation in females. It took advantage of highly polymorphic CAG repeats nearby HhaI sites in exon 1 of AR at the Xq13 region. X chromosome inactivation occurs early in embryogenesis and involves random inactivation and methylation of either one of the two X-chromosomes in all female cells. Once established, the X-chromosome inactivation pattern (XCIP) of a particular cell will be transferred to all of the progeny of that cell. As a result, a normal cell population of a healthy female usually shows a 50:50 ratio of inactivated paternal and inactivated maternal alleles. It is known that the malignant cells are derived from a single precursor cell, as all tumor cells contain the same inactivated X-chromosome, resulting in a monoclonal XCIP. The study on sensitivity demonstrates that clonal analysis by means of PCR offers a good method for studying the clonality in small DNA samples prepared from cryostat sections of tumors. This method could be applied to distinguish between benign and malignant (or premalignant) breast lesions, and thus be an adjunct diagnostic test in these morphologically ambiguous cases. Therefore, it would be very helpful in the design of rational prevention and treatment strategies.
MATERIALS AND METHODS
Patients and tissue samples
Tumor samples were obtained at the Tianjin Medical University Cancer Hospital between January 2000 and December 2005. Thirty cases of peri-PM, 30 cases of peri-PM with ADH, 30 cases of DCIS, 20 cases of developed canceration and 20 specimens of normal tissue were included in the study. The materials used in these studies were taken after pathologic diagnosis. The cases were all female patients with a median age of 48 years (range, 20~65 years). Multiple microdissections were performed on required cellular nests from different areas. More than 100 cells per case were obtained.
DNA extraction and restriction enzyme utilization
The genomic DNA was isolated using an EZNA DNA extraction kit. After determining the DNA concentration by ultraviolet spectrometry, the genomic DNA extracts (1 μg) were incubated at 37°C for 3 h in a reaction mixture of 20 μl (containing 2 μl of the accompanying buffer, 0.2 μl of 10 g/L BSA, and water) with or without 1 μl of Hha I. These samples then were heated to 65°C for 20 min to inactivate the enzymes in preparation for PCR amplification.
Polymerase chain reaction for AR analysis
Nested-PCR was used to amplify the androgen receptor gene fragment[9]. Each 10 μl of the digested DNA samples was added to a PCR reaction (final volume: 50 μl containing 20 pmol of each primer, 25 μl Premix Taq, and water) for amplification with the primer pair. The primer sequences used were (primers were offered by the Fourth Military University): AR1A: 5′-GAG GAG CTT TCC AGA ATC TG-3′; AR1B: 5′-CAT GGG CTT GGG GAG A-3′. An initial denaturing step of 7 min at 97°C was used to activate the Taq polymerase. We used 35 cycles starting with 94°C for 40 s, 56°C for 50 s, and 72°C for 1 min, ending with a 15-min extension step at 72°C. The PCR products (5 μl) were then seeded to the second reaction mixture with 2A and 2B as primers. Primer sequences were (primers were offered by the Fourth Military University): AR2A: 5′-TCC AGA ATC TGT TCC AGA GC-3′; AR2B: 5′-TGG GGA GAA CCA TCC TCA CC-3′.
PCR conditions were the same as described above.
Detection and scoring of clonality
The final PCR products of the androgen receptor gene were resolved in 20 g/L agarose gel to reveal whether they were informative or not. The samples were run on an 8% denaturing polyacrylamide gel to obtain better band separation. Samples that were restrictionenzyme digested then were compared with undigested samples. If both bands were present at similar intensities compared to the undigested lane, the case was scored as “polyclonal”. The cases were scored as “monoclonal” if one of the two bands disappeared after restriction enzyme digestion. The criterion for inclusion in the study was at least two separate assays showing the same result. The criterion for monoclonality was a qualitative significant decrease in the intensity of either the upper or lower allele compared with the intensity of the undigested control lane. The results were recorded, quantified and compared using a Gel Doc 1000 UV detection system and Multi Analyst software (Bio-Rad, Hercules, CA) for agarose gels.
RESULTS
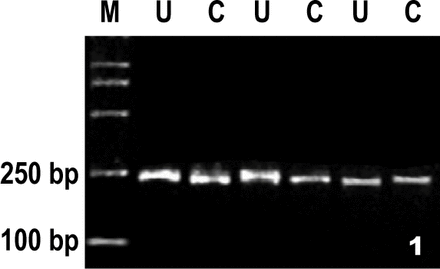
Determination of clonality was attempted on all samples from which sufficient DNA could be produced, including obvious malignancies, obvious cases of benign or reactive conditions, and atypical cases. The lane labeled “C”was cut with a restriction enzyme before PCR, the other lane in each pair labeled “U” was uncut. The lane “M” is DNA size Marker.
①The bands were visible at 230 bpDNA in 20 g/L agarose gel (Fig.1). This demonstrated sufficient DNA was produced.
②Nineteen specimens of the normal tissues were shown to be polyclonal.
③Twenty-four out of 26 (92.3%) cases of the peri-PM were shown to be polyclonal.
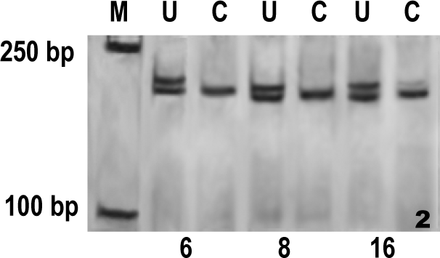
④Loss of polymorphism at the AR locus was revealed in all (27) the DCIS cases (Fig.2).
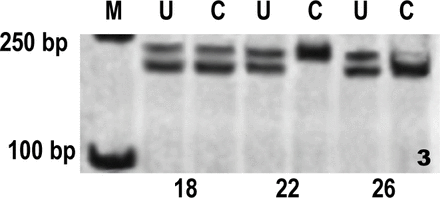
⑤Ten cases (10/25, 40.0%) of peri-PM with ADH indicated the monoclonality of the tumor (Fig.3).
⑥Among 16 cases of developed canceration, identical X chromosome inactivation (monoclonal alterations) was observed from both the peri-PM with ADH part and the DCIS portion in each case.
⑦The assay was repeated a number of times to prove the clonal status. Table 1 illustrates the assay for the determination of the clonality.
Resolving map of sufficient DNA produced in 20 g/L agarose gel. The bands were visible at 230 bpDNA in 20 g/L agarose gel.
Running map of DCIS on denaturing polyacrylamide gel. Case 6, 8 and 16 were scored as “monoclonal”. Case 6 and 8 showed one of the two bands disappeared after restriction enzyme digestion, whereas in case 16 a qualitative significant decrease displayed in the intensity of the upper allele.
Running map of peri-PM with ADH on denaturing polyacrylamide gel. Case 18 showed both bands were present at similar intensities to the undigested lane. Case 22 showed one of the two bands disappeared, whereas in case 26 a significant decrease appeared in the upper allele.
Data from outcome of the clonality assays.
DISCUSSION
Clonality analysis is potentially valuable in the identification of true neoplastic cells and prediction of clinical-biologic behavior. This would not only be useful to resolve morphologically difficult cases, but it could allow the process to become more objective. The most widely used technique for determining clonality is PCR of the human AR gene based on X-chromosome inactivation. The combination of microdissection and the image assay allows clonality analysis in virtually all tissue samples of females[13]. The method was applied in seborrheic keratoses, in which disappearance or significant reduction in intensity of one of two AR alleles was observed after HhaI digestion, but not in the normal control epidermis[14]. Clonality analysis was also performed in uterine leiomyomas[15], liver neoplasms[16], fibroepithelial tumors[17], apocrine adenosis of the breast[18], and pre-invasive breast diseases[19]. Although the AR assay method for detection of clonality is different, the principle is the same. Similarly, it appears that once sufficiently developed, this assay could have similar value in a wide range of tissues.
Our observations here revealed that all cases having a morphologic diagnosis of malignancy were monoclonal by the AR clonality assay. The majority of peri-PM were demonstrated to be polyclonal. This suggests that the risk of progression of these peri-PM lesions to invasive cancer is extremely low. In each case of developed canceration, both peri-PM with ADH and DCIS lesions showed the same monoclonal pattern, suggesting a clonal link. Among 25 cases of peri-PM with ADH, 10 cases indicated monoclonality of the tumor, as shown in 40% of the cases. These results contribute to the understanding of the genetic changes of peri-PM, and confirm the peri-PM with ADH as a precancerous lesion of the breast, which might be an important stage during the process of precursor developing to breast carcinoma. Two cases of peri-PM that were benign by morphology were shown to be monoclonal. There was surgical follow-up on them, one showing ductal hyperplasia; the other one is stable at present. It does carry significantly increased risk for a serious disease in the future. This outcome raises the question of the minimal clonal unit in breast tissue, and the earliest lesion that represents a clonal proliferation. Our work is consistent with these data because many benign cases in the current study were found to be polyclonal. In some cases different areas revealed distinct clonal status, which appeared that the specimen included a number of benign as well as malignant cells.
Further work is underway to increase the percentage of informative cases, and to decrease the complexity of the assay. We used laser-cut microdissection to gain the spindled cells, which allowed for rapid and accurate acquisition of cells in which we were interested[20,21], and tried to reduce the contamination of endothelial cells and fatty cells. We also compared the sensitivity of the polymerase chain reaction technique in the clonal analysis for detecting DNA in the paraffin-embedded tissues. No significant differences in the positive rates existed between general-PCR and nested-PCR. Thus the new method might be applicable and more convenient. Once the clonality analysis test is readily performed, it may be used in many difficult cases without a definitive morphologic diagnosis. It is likely that the method will form the basis of a revised system of classification in the future, with an underlying molecular genetic basis, while maintaining clinical relevance. In the longer term, this test has the potential for automation. One can conceive of its use for nonmorphologic-based diagnoses of clonality.
Acknowledgments
The authors gratefully thank Professor Qin Su of the TangDu Hospital, the Fourth Military University for fruitful discussions and for providing PCR primers.
Footnotes
This work was supported by a grant from the Tianjin Scientific Foundation Committee (No.033611911).
- Received January 10, 2007.
- Accepted November 12, 2007.
- Copyright © 2007 by Tianjin Medical University Cancer Institute & Hospital and Springer