Abstract
OBJECTIVE To screen differentially expressed genes in the development of human glioma and establish a primary molecular classification of glioma based on gene expression using cDNA microarrays.
METHODS Brain specimens were obtained from 18 patients with glioma, 10 males and 8 females, ages 14~62 with an average age of 44.4. The total RNAs of these glioma specimens and two specimens of donated brain of normal adults were extracted. BioStarH140S microarrays (including 8,347 old genes and 5,592 novel genes) were adopted and hybridized with probes which were prepared from the total RNAs. Differentially expressed genes between normal tissues and glioma tissues were assayed after scanning cDNA microarrays with ScanArray4000. Northern hybridization and in situ hybridization (ISH) were used to identify functions of novel genes. Those differentially expressed genes were studied with a Hierarchical method and molecular classification of glioma was preliminary carried out.
RESULTS Among the 13,939 target genes, there were 1,200 (8.61%) differentially expressed genes, of which 395 (2.83%) were novel genes. A total of 348 genes were up-regulated and 852 genes were down-regulated in the gliomas. The results of bioinformatical analysis, Northern hybridization and ISH revealed that those novel genes were highly associated with gliomas. There were multiple genes, such as the MAP gene cytoskeleton & matrix motility genes, etc, which were of relevance to classification by the Hierarchical method. Molecular classification of glioma using a Hierarchical cluster was in accordance with pathology and suggested a molecular process of tumorigenesis and development.
CONCLUSION Multiple genes play important roles in development of glioma. cDNA microarray technology is a powerful technique in screening for differentially expressed genes between two different kinds of tissues. Further analysis of gene expression and novel genes would be helpful to understand the molecular mechanism of glioma development.
keywords
Up to now, human glioma has been a disease incurable by neurosurgery and it is becoming a more and more serious health problem. Some difficult problems regarding human glioma such as rapid development, invasive ability and frequent recurrence have not been resolved.[1,2] Treatments including operations, chemical therapy, radiation and integrated therapy have not produced satisfactory results. Seeking other therapeutic approaches and elucidation of molecular pathologic mechanisms for glioma has become a goal in treating this cancer. Development of human glioma, involves complex variations of expression of multiple genes, which have been analyzed through microarrays.[3] Microarrays are useful in examining gene expression under differential conditions with various types of tissue, phases of development and gene alterations. [4] The analytical approach of microarrays, which includes K-mean clusters, artificial neural networks, and Hierarchical clusters, etc.[5] have been used to identify the function of genes related to human gliomas. These techniques organize the experimental samples on the basis of overall similarity in their gene-expression patterns. In our study DNA microarrays were used to profile changes in gene expression and differentially expressed genes were extracted in 18 cases of human gliomas. The molecular pathological difference between lower-grade gliomas and high-grade ones were analyzed through microarrays, combining clinical data and grades of pathology. This procedure provided evidence of invasion in human glioma and more target genes for glioma therapy.
Materials and Methods
Material
Tissue specimens
The 18 astrocytoma patient specimens used in this study were provided by department of Neurosurgery, Changzheng Hospital, the Second Military Medical University from July 2000 to May 2003. All the specimens were confirmed as astrocytoma by immunohistochemical targets (GFAP, NSE and S-100) and clinical pathology. The specimens, obtained during operations, were from 10 males and 8 females, ages 14 to 62 years with an average of 44.4 years. Pathology classification: 4 cases were astrocytoma grade I, 4 cases were astrocytoma grade II, 4 cases were astrocytoma III, 6 cases were glioblastoma, of which 2 cases of glioblastoma and one case of astrocytoma III was from a deceased patient. The 2 normal brain tissues were from donated specimens.
Material
BioStarH140s microarrays, including 14,112 spots which represented 13,939 genes (8,347 old genes and 5,592 novel genes) and 173 house-keeping spots, were provided by the United Gene Co. Hybridized cabins, silanized slides, silylated slide and silane slide were purchased from Sigma and TeleChem International, Sunnyvale, CA, USA.
Reagents and software
dNTPs were purchased from Promega(USA), protein enzyme K was from Sigma (USA); SuperScript II reversal transcription enzyme was from GIBCO/BRL. Image 3.0 software, PixPro 3.0.65 software, Cluster 2.11 software, Treeview software from: http://www. genome.standford.edu, NCBI database: http://www. ncbi.nlm.nih.gov; GCG database.
Experimental methods
Preparation probes
Total RNA was isolated from the 18 astrocytoma specimens and 2 normal brain tissues with an improved l-step method. The amount and quality of the total RNA were checked by GeneQuant (Clontech, Palo Alto, CA), electrophoresis on a 1.5% formamide agarose gel and by a test of heat stability in a 70 °C water bath. The fluorescent cDNA probes were prepared through reverse transcription of the isolated mRNAs with Superscript II enzyme and then 25 μg of total RNA was purified. [6] The RNA samples from healthy patients were labeled with Cy3-dUTP and those from cancerous patients with Cy5-dUTP. The two color probes were then mixed, precipitated with ethanol and dissolved in 20 μl of hybridization solution.
Hybridization, washing and drying
Fluorescent probe mixtures and chips were denatured at 95 °C for 5 min, and the denatured probe mixtures were applied onto the pre-hybridized chip under a cover glass. Chips were hybridized at 60 °C for 16 h.
The hybridized chips were then washed at 60 °C for 10 min each in solutions of 2 ⨯ SSC and 0.2% SDS, 0.1 ⨯ SSC and 0.2% SDS, and 0.1 ⨯ SSC, then dried at room temperature.
Detection and analysis
The chips were scanned with a ScanArray 4000 at 2 wavelengths to detect emission from both Cy3 and Cy5. The acquired images were analyzed using Image 3.0 software. Ratios of Cy5 to Cy3 were computed for each location on every microarray. Genes were identified as differentially expressed if the absolute value of the natural logarithm of the ratios was > 0.69. To minimize artifacts arising from low expression values, only genes with raw intensity values for both Cy3 and Cy5 of > 200 counts were chosen for differential analysis.
Hierarchical cluster
GeneMath 2.0 (Applied Maths Inc. USA)was adopted to analyze genes related to gliomas. The cluster results were exported with Treeview software and the ratio of genes on the microarrays were analyzed. We analyzed the relativity of gene expression in the 18 different grades of human glioma and put patients with similar expression into border genus, and performed a molecular pathological classification of different grades of astrocytoma.
Northern blots
Equal amounts of total RNA (30 µg) extracted from normal brain tissues and 7 glioma tissues were loaded onto 1.5% gels. The blots were scanned with a Cyclone instrument and the data were analyzed with Parkard 3.0 software. The 5 genes relating to astrocytoma were selected for preparation probes, which were marked with a-32P dCTP. The experiment was operated according to Northern blot routines.
In situ hybridization
1) Preparation of tissue slices: Frozen normal brain tissues and adult glioma tissues were sliced at 8 µm in a sagittal plane and mounted onto Probe On slides purchased from Fisher.
2) Design specific primer: the LNX gene was amplified by PCR with a pair of specific primers (Fl :5 ‘-taatacgactcactataggggaaggcgcttctgttgctggtcttgcc-3 ‘, F2: 5’-attaaccctcactaaagggaaaagcttcatctcggggtc-tg-taggc-3’ ). Anti-sense RNA (cRNA) / sense (mRNA) was transcribed by T3/T7 RNA polymerase in the presence of labeled DIG-UTP nucleotides.
3) Hybridization and display: After removal of the template by incubation with DNAse I, RNA transcripts were precipitated in ethanol containing LiCl, and dissolved in DEPC treated water. Sections were fixed for 5 min with 4% formaldehyde. To reduce non-specific binding of the probe, we included an acetylation step into the prehybridization phase. After 16 h of hybridization at 49°C, samples were rinsed with a SSC solution of increasing stringency at constant temperature (49°C ). The DIG-labeled probe was visualized anti-DIG antibody and subsequent alkaline phosphatase staining according to the protocol supplied by the producer. The concentration of anti-DIG antibody was 150 mU/ml.
Results
RNA hot-stabilization experiment
The RNAs of normal brain tissues and glioma tissues were examined through hot-stabilization experiments. The results revealed that strips of RNA were pure and consistent in experiments.
Scanning and analyzing microarrays
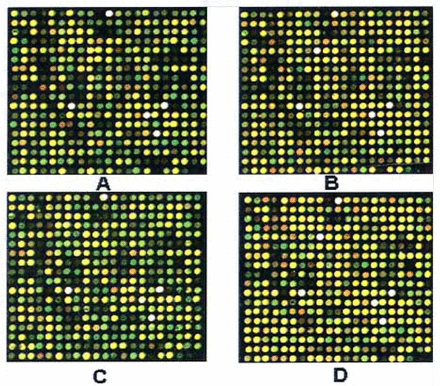
The hybridization signals were determined after the probes were hybridized in the microarray. Of all the 13,939 human genes analyzed through microarray experiments, a total of 1,200 genes (8.61%) revealed different expression in more than 50% of the individual patient specimens using a fold ratio of < 0.5 or > 2 as criteria for cut-off (Fig. 1). There were 395 novel genes not accepted in the GeneBank, 348 genes were up-regulated and 852 genes were down-regulated. Bioinformatic analysis identified 124 genes related to genes for cell signal transduction, 74 associated with cellular metabolism genes, 43 were cytoskeleton & motility genes, 21 were oncogenes and tumor suppressor genes, 48 for synthesizing protein, 31 for immunity, 24 for developmental processes, 21 genes for ion-channels, 21 for DNA combination, 17 for the cell cycle, 13 for cell receptors, 4 for cell apoptosis, and 5 for DNA repair and restructaring. It showed that there were multiple genes associated with human glioma.
Gene expression of differential grades of human glioma tissues: The green spots represent down-regulated genes in human gliomas, the red spots represent up-regulated genes in tumors and the yellow spots represent similar expression between human glioma and normal brain tissue; A, B, C D represent gene expression of astrocytoma I, II, III and IV.
Results of Hierarchical cluster
A total of 1,200 differentially expressed genes were analyzed with a Hierarchical cluster. The gene expression profile of human gliomas from 7 living patients No. 2,3,14,16,17,4,15 were similar, the pathology analysis showed all of these 7 cases were astrocytoma I and II. With further classification, we found the gene profiles of patient No. 2 and 3, No. 14 and 16, No. 15 and 7, No. 17 and 4 were similar, respectively, and suggesting tumor cells have similar molecular biological actions. In the highly malignant group, the gene expression of No. 5, 6 and 9 were similar and No.18, 19, 10, 13, 8, 11 were similar, which revealed that the extent of malignantcy was closely related to expression profiles (Fig.2). Therefore, gene expression classification could distinguish consistent with clinical pathology classification.
Molecular pathological classification of 18 gliomas based on gene expression: Every vertical line represents gene expression of a human glioma and a rank line represents expression of every gene; the red diamonds represent up-regulation, the green ones represent down-regulation, the white ones represent lowly expression; the right of the Fig. represents a number of the gene clone and GenBank, and where blank there is a novel gene.
Northern-blots
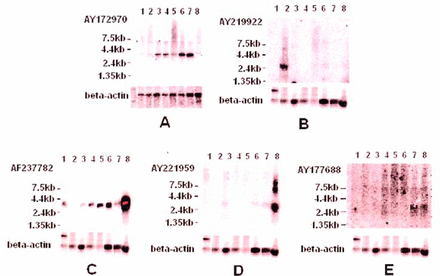
Five novel genes (AY 172970, AY219922, AF237782, AY177688, AY221959)differentially expressed in glioma on microarrays were examined by Northernblots. Compared to the normal brain tissue, AY 172970 was significantly up-regulated in gliomas as seen in the hybridization signal in Fig.3A, while AY219922, AF237782, AY221959, AY177688 were significantly down-regulated in the gliomas in Fig.3B, C, D, E. In addition, AY177688 was slightly up-regulated in recurrencent glioblastoma compared to normal brain tissue. The results of Northern-blots were identical to the results of microarrays.
Expression of 5 novel genes in normal brain tissue and human glioma: A Expression of the AY172970 gene in normal brain and 7 gliomas 1. Recrudescent glioblastoma IV; 2~4. Glioblastoma IV; 5. Astrocytoma HI; 6. Astrocytoma II;7. Astrocytoma I;8.Normal brain tissue; B Expression of the AY219922 gene in embryonic brain, normal brain and 6 gliomas 1. Embryonic brain; 2. Normal brain tissue; 3. Astrocytoma I; 4. Astrocytoma II; 5. Astrocytoma III; 6,7 Glioblastoma IV; 8. Recrudescent glioblastoma IV; C.D.E 3 hybrizations on the same film 1. Astrocytoma I; 2. Astrocytoma II; 3. Astrocytoma III; 4~6. Glioblastoma IV;7. Recrudesce glioblastoma IV;8. Normal brain tissue; AF237782, AY221959, AY177688 novel genes were down-regulated in differential grades of gliomas, while AY177688 was slightly up-regulated in recrudesce glioblastoma IV.
Results of in situ hybridization
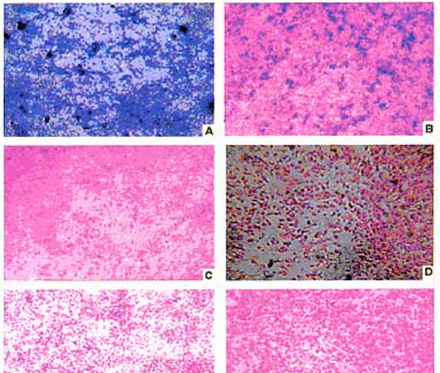
In situ hybridization studies were performed to determine the expression patterns of genes in gliomas and normal adult brain. Our results showed AF237782 expressed in the cytoplasm in the normal brain tissue (Fig.4), however, its expression decreased remarkablely in the glioma cells, even absent in glioblastoma cells. Most of the in situ hybridization analysis results agreed with the microarrays and Northern blot data. Taken together, these studies indicated that AF237782 was of good diagnostic value for glioma patients.
Expression of AF237782 novel gene in brain and human glioma (NBT/ BCIP) ⨯ 200: A normal brain tissue; B Astrocytoma I; C Astrocytoma II; D Astrocytoma III; E, F Glioblastoma IV.
Discussion
Pathological classification of a tumor is one of most credible indexes that can reflect clinical characteristics of the tumor. Although some of the same grade gliomas have been treated identically in clinical studies, significant differences in prognosis and therapy exist. The differences in expression of some genes related to invasion and recurrence in the same grades of glioma have been identified,[7,8] while clinical pathological classification can not reflect this difference. The biological characteristics of glioma are based on differences in gene expression. It is of importance that we search for tumor marker genes, which can be used to estimate invasion, proliferation, differentiation, and prognosis of human gliomas reliablely and conveniently.
Detecting key novel genes in human glioma
Gene expression in different grades of glioma was analyzed in depth through microarrays, which can be used to analyze expression of both novel and known genes in the development of glioma. Microarrays possess significant merits compared to other methods and can produce comparisons between novel genes and human glioma. Therefore, we focused on these important genes. Five genes which possess important functions were isolated to distinguish tumor stages and tumor subtype. AY172970 is a novel BEACH protein which can accelerate tumor cell division and participate in regulating human glioma signal transduction. [9] Northern blots confirmed that it was up-regulated in lower grades of glioma and down-regulated in high grades of glioma compared to lower ones, suggesting that it accelerates tumor cell division in the primary stages of glioma and could be a potential index to estimate malignancy of gliomas. AF237782 is another novel human numb protein which participated in notch approach and TGF-β signal transduction.[10] It is a potential suppressor gene which restrains development of glioma through the above mechanism. AY219922 is a novel human ARM protein which participates in signal transduction and is related to APC and β-catenin. [11,2] The down-regulation of AY219922 is associated with the development of human glioma. AY177688 is a novel J-Domain protein which participates in dopamine signal transduction, as a member of a molecular partner family.™ The AY221959 protein contains a novel BTB/POZ domain, involved in protein reciprocity and transduction. All of these novel genes provide new information about the molecular mechanism of glioma formation.
Molecular classification based on gene expression
It is well-known that the biological characteristics of glioma are decided by gene clusters rather than by individual genes. Gene clusters can reflect the development of glioma more precisely than individual genes. Pomeroy et al. [14] identified multiple genes related to classification in medulloblastoma by cDNA microarrays, suggesting that molecular pathological classification based on gene expression reflects a tumor’s nature more directly than morphological classification. Golub et al.[15] established an approach to identify molecular differences between AML and ALL based on gene expression through microarrays, which could be used to classify leucocythemias and predict its prognosis. Eisen et al. [16] established a mathematical model of gene clusters and confirmed that gene expression related closely to the biologic nature of the tumors. Alizadeh et al.[17,8] found many kinds of genes could be used to diagnose and produce a prognosis for a malignant tumor, considering that molecular pathological classification based on gene expression could explain clinical phenomena which traditional classifications could not explain. These studies contribute to identification of the essential character of glioma, and thereby distinguish tumors which have similar morphology. We analyzed the characteristics of gene expression in 18 glioma specimens combined with clinical data, and found comparable gene expression in the same grades of glioma. There was a significant difference between lower and higher grades of glioma. Our results based on Hierarchical clusters revealed that the MAP gene, cytoskeleton and matrix motility genes were significantly related to molecular classification. The MAP gene is a major cytoskeleton component associated with microtube assembling and is useful for neuroglioma diagnosis and p53 regulation. [19] The super-family of adhesion proteins and cellular matrix genes participate in adherence and invasion, indicating molecular pathological classification was in accordance with cell morphology. Expression and regulation of genes determine cell morphology, behavior and function, and further clinical behavior. The details of gene alteration can be useful for understanding development and prognosis of human glioma.
cDNA microarrays can be employed to screen differentially expressed genes high-throughout and high-efficiency to study tumorgenesis. Because of cost, we examined only 18 cases of glioma specimens and analyzed the relationship between gene classification and pathological classification by cDNA microarrays. Furthermore, we shall confirm reliability of novel genes combined with the results of clinical data, and seek for principles of gene expression. At the present time, it is very important to improve clinical therapy by making use of results from basic studies. It would be helpful for clinical diagnosis, therapy and development of prognosis to understand the molecular mechanism of glioma carcinogensis by using microarray technology.
Footnotes
This work was supported by the National Natural Science Foundation (30471777) and the Natural Science Foundation of Shanghai (034119835).
- Received August 4, 2004.
- Accepted December 10, 2004.
- Copyright © 2005 by Tianjin Medical University Cancer Institute & Hospital and Springer