Abstract
OBJECTIVE To quantitatively explore the expression of Syndecan-1 and heparanase-1 in esophageal cancer tissue as well as their relationship with the clinicopathological factors, in order to evaluate their roles in tumor invasion and metastasis.
METHODS Real-time fluorescence quantitative PCR (Q-PCR) was used to analyze the expression levels of Syndecan-1 and heparanase-1 genes, participants included 67 cases with esophageal cancers and 32 healthy volunteers.
RESULTS The expression of Heparanase-1 gene in esophageal cancers was higher than that in normal esophageal tissue (P < 0.001), and the expression of Syndecan-1 gene in the normal esophageal tissue was higher compared with esophageal cancers (P < 0.001). The positive rates of Syndecan-1 and Heparanase-1 gene in esophageal cancer were 13.4% (9/67) and 85.1% (57/67). The expression of Syndecan-1 and Heparanase-1 genes was significantly related to differentiation, depth of infiltration, lymph node metastasis, vessel metastasis, and TNM stages of disease (P < 0.05). In an attempt to measure the association between the 2 agents, this study found that the expression of Syndecan-1 mRNA had a significantly negative correlation with the expression of Heparanase-1 mRNA by using Spearman rank correlation test (OR = -0.572, P < 0.001).
CONCLUSION Syndecan-1 and Heparanase-1 play important roles in the invasion and metastasis of esophageal cancer. The reduction of Syndecan-1 and/or the increase of Heparanase-1 may influence the invasion and metastasis of malignant tumors. Thus the combination assay of Syndecan-1 and Heparanase-1 may contribute to the diagnosis and treatment of malignant tumors.
keywords
Introduction
Esophageal cancer remains one of the most aggressive malignant carcinomas and its mortality has gradually increased in recent years. The main cause of death among patients with esophageal cancer is invasion and metastasis. Many processes including adherence, degradation, motility, and angiogenesis are involved in the invasion and metastasis of esophageal cancer[1]. In the process of invasion and metastasis, tumor cells must break the barrier of the extracelluar matrix (ECM) and the basement membrane (BM). Syndecan-1 is a set of transmembrane heparin sulfate glycoproteins (HSPGs), which is the main component of the barrier. Heparanase-1, an endo-β-D-glucuromdase, is one of the enzymes involved in the ECM remodeling process, and it can degrade heparin (HS) side chains of heparin sulfate proteoglycans (HSPG), such as Syndecan-1. Some authors have found that the loss of Syndecan-1 in some carcinomas can enable the invasion and metastasis of malignant cells[2,3]. Heparanase-1 up regulation has been documented in an increasing number of primary human tumors[4,5], including pancreas[6], bladder[7], gastric[8], cervical[9], and colorectal[10] tumors. However, there are few reports about the expression of Syndecan-1 and Heparanase-1 in esophageal cancer. This study is to quantitatively explore the expression of Syndecan-1 and Heparanase-1 in human esophageal cancer by real-time fluorescence quantitative PCR.
Patients and Methods
Patients
Between Jan 2008 and May 2009, 67 patients with esophageal cancers who underwent esophagectomy with lymph node dissection in the Department of Gastrointestinal Surgery at Qingdao Municipal Hospital were included in this study. None of the patients had received chemotherapy or radiotherapy. The dissected specimens were histologically examined by 2 experienced pathologists who were blind to the data and results of other diagnostic tests. The clinicopathologic characteristics were evaluated according to the guideline of Union for International Cancer Control (UICC). Thirty-two samples of normal esophageal tissue were obtained from the healthy individuals undergoing endoscopies to use as the controls. For the Real-time fluorescence quantitative PCR study, tissue samples were kept at -80°C immediately after surgical removal.
Cell culture
EC9706, a kind of esophageal cancer cell line, was obtained from Chinese Medical University. EC9706 cells were plated in a 6-well plate at 5 × 104 cells/L. And the cell lines were propagated in RPMI1640 with 10% FCS, 100 unit/mL penicillin, and 100 mg/mL streptomycin at 37°C in humidified air with 5% CO2.
Reverse transcription-PCR
Total RNA was isolated from fresh tissue and cell lines by using TRIZOL Reagent (TaKaRa Biotechnology [Dalian] Co., Ltd.). The integrity of RNA was checked electrophoretically and quantified spectrophotometrically. Reverse transcription condition was performed at 43°C for 45 min, 95°C for 5 min and 0°C for 5 min. The PCR reactions were performed with specific primers according to the condition: 94°C for 5 min, 94°C for 30 s, 60°C for 1 min, 72°C for 1 min, 72°C for 10 min. All amplification reactions were carried out in a final volume of 25 μL containing 2.5 μL 10 × PCR Buffer, 0.5 μL dNTPMix, 0.8 μL PCR Forward Primer, 0.8 μL PCR Reverse Primer, 0.5 μL TaqMix DNA polymerases, 2.5 μL RT product (Qingdao ALP Biotech Co., Ltd.). The following primers were used for amplification: i) Heparanase-1, forward primer 5′-GAA AGA CGG CTA AGA TGC TGA AGA-3′ and reverse primer 5′-GGT AGC AGT CCG TCC GTC CAT TCA AA-3′ (PCR Prouduction, 101 bp); ii) Syndecan-1 forward 5′-ATG GAG GTC CTT CTG CCA CC-3′ reverse, 5′-TCT CCC CCG AGG TTT CAA AG-3′ (PCR Production, 111 bp); iii) GAPDH, 5′-TCA TGG GTG TGA ACC ATG AGA A-3′ and reverse primer 5′-GGC ATG GAC TGT GGT CAT GAG-3′ (PCR production, 146 bp). Same amount of the amplification products were run on 2% agarose gel stained with ethidium bromide and was observed under a UV transilluminator.
Real-time fluorescence quantitative PCR
Reverse transcription and PCR amplification were performed based on manufacturer’s recommendations (SYBR Prime Script RT-PCR Kit, TaKaRa Biotechnology (Dalian) Co., Ltd.). The total RNA was heated to 37 °C in a bath for 25 min for reverse transcription, and the reverse transcriptase was inactivated in a bath at 85°C for 1 min. Complementary DNA (cDNA) was synthesized from 1 μg of total RNA in a 10 μL reaction mixture containing 2 μL 5 × PrimeScriptTM Buffer, 0.5 μL PrimeScriptTM Enzyme Mix I, 0.5 μL Oligo DT Primer, 0.5 μL random 6-mers and 1.8 μg RNA. The remainder was ddH2O.
The real-time fluorescence quantitative PCR analysis was performed on an automated sequence detection system (Rotor-Gene 3000 Real-Time PCR System). PCR amplification with the specific primer was performed in 20 μL final reaction volumes containing 10 μL SYBR Primer EX TaqTM II, 0.8 μL PCR Forward Primer, 0.8 μL PCR Reverse Primer, 2 μL cDNA, 6.4 μL dH2O. The initial denaturation step of the PCR reaction was for 30 s at 95°C, and then the PCR reaction was run for 40 cycles under the following conditions: denaturation for 5 s at 95°C and annealing for 5 s at 95°C. The following primers were used for amplification: i) heparanase-1, forward primer 5′-CCT TGC CAC CTT TAA TGG AA-3′ and reverse primer 5′-AAG CAG CAA CTT TGG CAT TT-3′ (PCR production, 105 bp); ii) Syndecan-1, 5′- GGG ACT CAG CCT TCA GAC AG -3′ and reverse primer 5′- CTC GTC AAT TTC CAG GAG GA -3′ (PCR production, 128 bp). GAPDH transcripts were monitored as a control to quantify the transcripts of the genes in each sample. The primer sequence of GAPDH in real-time fluorescence quantitative PCR was the same as the sequence used in reverse transcription-PCR.
The relative mRNA levels were expressed as Ct values, where Ct is defined as the threshold PCR cycle at which the amplification product was first detected, and ΔCt is the difference between the Ct values of Heparanse-1 and GAPDH, Syndecan-1 and GAPDH. Normal esophageal tissue cDNA was used as a calibrator to generate a ΔΔCt value (ΔCt test sample-ΔCt normal sample). The N-fold differential expression in heparanase-1 compared with calibrator was expressed as 2-(ΔΔCt). To verify the sequence of products, regular PCR was performed and ran the PCR products on a 2% agarose gel in parallel with Marker I before staining with ethidium bromide. The authenticity of PCR products was confirmed using a BLAST search of the sequence through National Center for Biotechnology Information.
Statistical analysis
Statistical analysis was performed by employing the Statistical Package for Social Sciences software program version 16.0 (SPSS Inc, Chicago, Illinois, USA). χ2 test and 2-sample t-test were used to compare the demographic and clinical data between groups. The expression levels of Syndecan-1 and Heparanase-1 in esophageal cancer and normal esophageal tissue were expressed as Mean ± SD. The correlation between relative Syndecan-1 mRNA and heparanase-1 mRNA in esophageal cancer was evaluated statistically using the Spearman rank correlation test. A P-value of less than 0.05 was considered statistically significant.
Results
Heparanase-1 mRNA expression and clinicopathologic factors
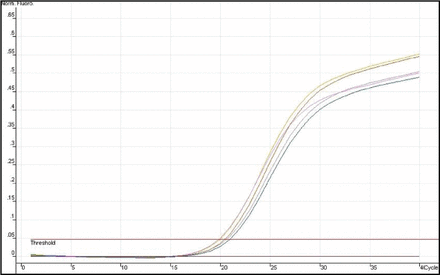
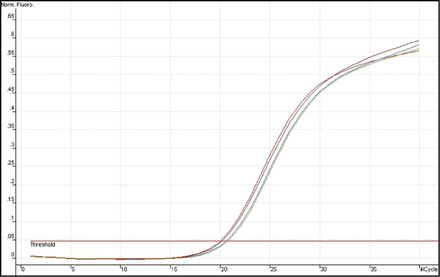
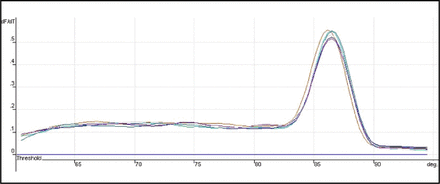
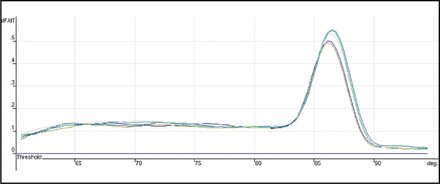
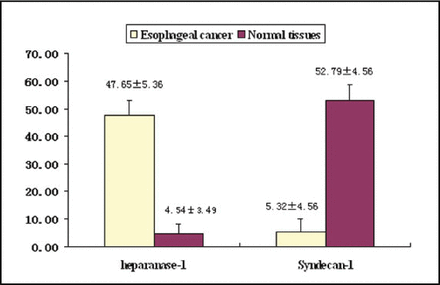
The expression of Heparanase-1 mRNA and Syndecan-1 mRNA was detected quantitatively by using Real-time fluorescence quantitative PCR. The amplification plots showed the typical “S” (Figs. 1 and 2), and the melting temperature profiles of amplification were determined to show the specificity of amplification (Figs. 3 and, 4). Representative RT-PCR results are shown in Fig. 5 and Fig. 6. In esophageal cancer cell line EC9706, the test could always detect the expression of Heparanase-1, while Syndecan-1 expression was never detected. In the 32 normal esophageal tissue, the test could only detect 4 cases of Heparanase-1 mRNA and 29 cases of Syndecan-1 mRNA. In the 67 cases of esophageal cancer, 57 cases had positive Heparanase-1 expression and 9 cases had positive Syndecan-1 expression. As shown in Fig. 7, the expression of Heparanase-1 mRNA in esophageal cancer and normal esophageal tissue was (47.56 ± 5.36) and (4.54 ± 3.49), while the expression of Syndecan-1 in normal esophageal tissue (52.79 ± 4.56) was significantly higher than that in esophageal cancer tissue (5.32 ± 4.56) (P < 0.001).
Representative results of RT-PCR amplification plots for Heparanase-1mRNA.
Representative results of RT-PCR amplification plots for Syndecan-1mRNA.
Representative results of RT-PCR Dissociation plots for Heparanase-1 mRNA.
Representative results of RT-PCR Dissociation plots for Syndecan-1 mRNA.
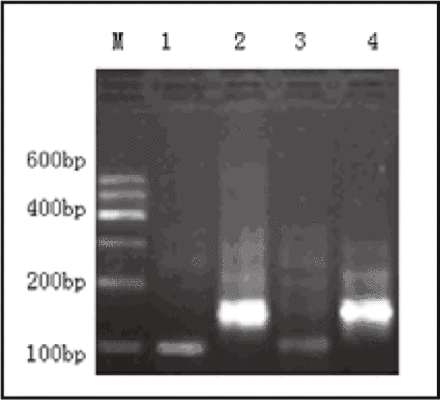
Agarose gel electrophoresis of the Heparanase-1 PCR products in esophageal cancers. Lane M: Marker 1 (600 bp); Lane 1, 3: The expression of Heparanase-1 mRNA (101 bp) in the esophageal cancers; Lane 2, 4: The expression of GAPDH mRNA (146 bp) in the esophageal cancers.
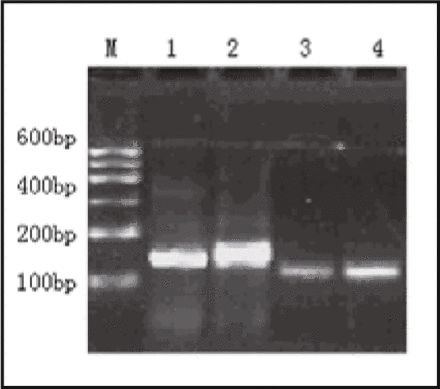
Agarose gel electrophoresis of the Syndecan-1 PCR products in normal esophageal tissue. Lane M: Marker 1 (600 bp); Lane 1, 2: The expression of GAPDH mRNA (146 bp) in normal esophageal tissue; Lane 3, 4: The expression of Syndecan-1 mRNA (111 bp) in normal esophageal tissue.
The expression of Syndecan-1 and Heparanase-1.
Expression and clinicopathological factors of syndecan-1 and heparanase-1 in esophageal cancer
The correlation between Heparanase-1 and Syndecan-1 and clinicopathological factors in the 67 esophageal cancer specimens was analyzed statistically as described in “Statistical Analysis”. As shown in Table 1, the differentiation grade of carcinomas was either well differentiated n = 13 (68.4%), or moderated n = 44 (91.7%). The expression rate in advanced carcinomas at stages T3-T4 was significantly higher than in the earlier T1-T2 stages (P < 0.05). Forty-five patients (91.8%) presented lymph node metastasis. Concerning hematogenous metastasis, 43 patients (91.5%) presented this metastasis, whereas in 14 patients (70.0%) it was absent. Fourteen patients (63.6%) were at initial stags (I or II), whereas, 43 patients (95.6%) were at advanced stages (III or IV) on UICC. As shown in Table 1, Syndecan-1 activation was correlated with differentiation, depth of infiltration, lymph node metastasis, hematogenous metastasis and TNM stages of disease (P < 0.05).
Expression and clinicopathological factors of Syndecan-1 and Heparanase-1 in esophageal cancer.
Correlation between Syndecan-1 mRNA and Heparanase-1 mRNA in esophageal cancer
By using the Spearman rank correlation test, it was found that the expression of Syndecan-1 mRNA had significant negative correlation with the expression of Heparanase-1 mRNA (OR = -0.572, P < 0.001, Table 2).
Correlation between Syndecan-1 mRNA and Heparanase-1 mRNA in esophageal cancer.
Discussion
Invasion and metastasis are the main causes of death due to malignant tumors. Thus, how to block the invasion and metastasis of the carcinoma timely and effectively has become the dilemma of the cancer research world. The critical process of the invasion and metastasis of carcinoma is to break the barrier which is composed of ECM and BM. The barrier mainly contains structural proteins and glycosaminoglycans (GAGs). In the past decades, people mostly focused on the properties of some enzymes, such as MMPs which use structural protein as a substrate. However, the 2 critical agents, Syndecan-1 and Heparanase-1, which play a more important role in the invasion and metastasis, were so far neglected. Syndecan-1 is one member of the family of cell surface transmembrane heparan-sulfate proteoglycans (HSPGs) which are the major component of the GAGs, and Heparanase-1 is the exclusive endoglycosidase which can degrade the hepran-sulfate side (HS) chains of HPSGs.
Syndecan-1, as the most exclusively studied member of the HPSGs, constitutively expressed on the surface of many mature epithelial cells but not on mesenchymal cells. It consists of a core protein to which side chains of complex glycosaminoglycan heparin sulfate (HS) are attached. A C-terminal cytoplasmic domain and an N-terminal extracellular domain, covalently attached to long unbranched carbohydrate polymers, called glycosaminoglycans (GAGs)[11,12]. Syndecan-1 functions in cell-cell and cell-extracellular matrix interactions via its heparin sulphate chains[13]. The extracellular domain of Syndecan-1 can inhibit cell invasion[14-16] and suppress malignant growth in different carcinoma cell lines[17]. Syndecan-1 expressed at the cell surface is typically down-regulated in metastatic phenotypes.
Heparanase-1 is an endo-β-D-glucuromdase that can degrade heparin sulfate and is implicated in a variety of biological processes. Over-expression of Heparanase-1 was found in various kinds of malignant tumors, but expression was either absent or low in normal tissue[18-22]. The overexpression of Heparanase-1 mRNA has been reported to correlate with the metastatic or invasive potential of tumor cells in vitro and in vivo[23,24] and with poor patient prognosis[25].
In regard to Syndecan-1 and Heparanase-1, many authors have already reported their relationship with the colonic carcinoma, gastric carcinoma, gallbladder carcinoma, etc. However, to date, there are very few reports on detecting the expression of Syndecan-1 and Heparanase-1 in esophageal cancer, especially the combination of the two. Meanwhile, most of the methods for detecting the expression of Syndecan-1 or Heparanase-1 in carcinoma were immunohistochemistry and cytology which have lower sensitivity and specificity than Real-time fluorescence quantitative PCR. In this study, Real-time fluorescence quantitative PCR was used to quantitatively detect the expression of Syndecan-1 mRNA and Heparanase-1 mRNA in esophageal cancer.
In the present study, the positive rate of Syndecan-1 mRNA expression in esophageal cancer (13.4%) was significantly lower than that of normal esophageal tissue (90.6%) (P < 0.05). The expression of Syndecan-1 mRNA was closely related to the clinic pathological factors of carcinomas. With the depth of infiltration increasing, the detection rate of Syndecan-1 gradually decreases: 4 cases were Syndean-1 positive in 47 patients of lymphatic metastasis, while 5 cases were positive in 20 patients without lymphatic metastasis. The difference was statistically significant (P < 0.05). This study also demonstrated that the expression of Syndecan-1 mRNA has a significant negative correlation with hematogenous metastasis, differentiation grade and TNM stages. The results indicated that the loss of Syndecan-1 mRNA is closely related with the clinicopatholigical factors that reflect the invasion and metastasis potential and prognosis. Thus, a conclusion can be drawn that the loss of Syndecan-1 plays an important role in the carcinoma generating, development, invasion and metastasis. JIRO[26] previously demonstrated that the expression of syndecan-1 is significantly lower in gastric type than in ordinary type and complete-intestinal type cancers. Wiksten et al.[27] reported that abatement or loss of syndecan-1 expression is highly correlated to the lymphatic metastasis, invasion depth, TNM stages and prognosis of gastric carcinoma.
In this study, it was found that the positive rate of Heparanase-1 in esophageal cancer is 85.1%, significantly higher than that in normal esophageal tissue 12.5% (P < 0.05). Heparanase-1 mRNA tended to express more in those patients with lymphatic metastasis, hematogenous metastasis, serosal invasion, advanced stage of disease, and moderated tumors. These results supported the findings to date from Wu[28] and EI-Assal[29]. That Heparanase-1 mRNA expression statistically related with these factors implies that the invasion and metastasis induced by Heparanse-1 might indicate a poor prognosis for esophageal cancers. Over-expression of Heparanase-1 was thought to contribute to tumor invasion, metastasis[30], and inflammatory reactions[31], and the enzyme was moderately up-regulated especially in metastatic cancers[32].
By this study, it could be found that the expression of Heparanase-1 mRNA had a significantly negative correlation with the Syndecan-1 mRNA by the Spearman rank correlation test. The difference was statistically significant (OR = -0.572, P < 0.001). It was also demonstrated that expression of Syndecan-1 on melanoma cell surface and its degradation by Heparanase-1 are important factors in the control of tumor cell invasion and metastasis[33]. So far, there have been very few reports on the relationship between the Syndecan-1 and Heparanase-1 genes. The combination assay of the two plays an important role in cancer diagnosis.
In conclusion, our study showed that Syndecan-1 and Heparanase-1 play important roles in the growth, invasion, and metastasis of esophageal cancer. It also showed that the reduction of Syndecan-1 and the increase of Heparanase-1 may accelerate the invasion and metastasis of esophageal cancer. Limited by sample size, further investigations with larger sample sizes will be necessary to confirm the expression of Syndecan-1 and Heparanase-1 in esophageal cancer. Further studies on the significance of Syndecan-1 and Heparanase-1 to the invasion and metastasis in esophageal cancer are also necessary.
Conflict of interest statement
No potential conflicts of interest were disclosed.
- Received July 12, 2010.
- Accepted August 10, 2010.
- Copyright © 2010 by Tianjin Medical University Cancer Institute & Hospital and Springer