High cancer heterogeneity and aggressiveness are major factors contributing to the poor prognosis of patients with advanced cancer1–5. Early detection of cancer can significantly improve the prognosis of patients with early-stage cancer after standard treatment. Although traditional cancer detection technologies, such as endoscopy for gastrointestinal cancer, mammography for breast cancer, HPV DNA detection for cervical cancer, and computed tomography and positron emission tomography-computed tomography, are effective, these largely non-invasive screening methods pose various challenges, including patient resistance due to discomfort during the procedures, high costs, and requirements for specialized equipment and experienced physicians. Consequently, conducting these procedures on a large scale is difficult.
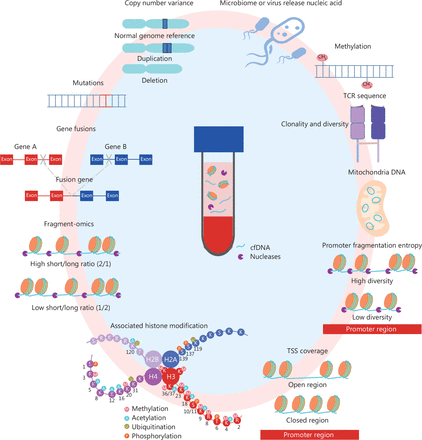
The emergence of circulating cell-free DNA (cfDNA), newly identified non-encapsulated DNA molecules, has revolutionized cancer early detection by offering a comprehensive genomic perspective. The abundant information contained in cfDNA includes mutations, copy number variations, gene fusions, DNA methylation, microbiome or virus-released nucleic acids, T cell receptor sequences, mitochondrial DNA abnormalities, fragment ratios, associated histone modifications, transcription start site (TSS) coverage, and promoter fragmentation entropy (PFE)6,7. (Figure 1). These innovative technologies have shown potential in accurate and practical approaches to large-scale cancer screening and preoperative evaluation.
cfDNA, a new non-invasive technology for early cancer detection. The multi-omics approach—which includes mutations, copy number variations (CNVs), gene fusions, DNA methylation, microbiome or virus-released nucleic acids, T cell receptor sequences, mitochondrial DNA abnormalities, fragment ratio, associated histone modifications, TSS coverage, and promoter PFE—can be effectively used for early cancer detection.
Although cfDNA-based technologies offer substantial promise for early cancer detection, they also have several limitations. For instance, clonal hematopoiesis leads to most cfDNA mutations, which can be confused with the mutations induced by the circulating tumor DNA, thereby affecting the accuracy of cancer diagnosis and the method’s clinical utility. Additionally, mutated gene panels have low detection rates in patients with early-stage cancer8, and DNA methylation, although stable, cannot reflect cell-specific programs or molecular dynamics9. Moreover, the bisulfite processing of cfDNA results in degradation of at least 84%–96% of the DNA, thus leading to a loss of valuable information10. Consequently, samples of hundreds of nanograms (ng) of DNA (derived from 10 mL blood) are required to mitigate the effects of clonal hematopoiesis, rare counts in cfDNA mutations, and the substantial amount of degraded DNA in cfDNA methylation analyses.
Although the limitations of some cfDNA-based technologies, including fragment ratio, associated histone modifications, TSS coverage, and PFE, are unclear, these signatures can be used to infer gene expression levels and chromatin accessibility status, thereby indicating the functional state of cancer cells, and providing comprehensive information on dynamic biological processes and heterogeneity during tumorigenesis, in contrast to approaches focusing on certain mutations and CpG islands. Hence, the enormous potential of these cfDNA-based technologies, including fragment ratio, associated histone modifications, TSS coverage, and PFE, warrant further in-depth exploration.
The cfDNA fragment ratio reveals abnormalities in disease
Most cfDNA in healthy individuals originates from myeloid and lymphocyte cells, which have similar chromatin accessibility and nucleosome footprints. Therefore, the cfDNA fragment ratio is stable in healthy individuals, with a median correlation of fragment ratio of 0.9911. However, in individuals with cancer, tumor-derived cfDNA can disrupt this balance, thus decreasing the median correlation of fragment ratios to 0.84. The cfDNA fragment ratio features in breast, colorectal, lung, ovarian, pancreatic, gastric, and bile duct cancer have been found to achieve sensitivities ranging from 57% to > 99% and 98% specificity, with an overall area under curve (AUC) value of 0.94. Recently, multi-center clinical trials have surveyed cfDNA fragment ratios in 724 individuals from the US, EU, or Hong Kong (China) with hepatocellular carcinoma, or individuals with average or high risk for hepatocellular carcinoma. In the average-risk population, a sensitivity of 88% and specificity of 98% for detecting cancer were observed, whereas the high-risk group had a sensitivity of 85% and specificity of 80%12. These studies demonstrated the potential clinical application of cfDNA fragment ratio in cancer early detection. Furthermore, the detection of abnormal fragment ratios in target genome regions would be suitable for clinical application on a large scale.
cfDNA associated histone modifications indicate actively transcribed genes
Histone modifications play crucial roles in regulating chromatin structure and the activity of RNA. Different combinations of these modifications serve as distinct markers of gene promoter regions, enhancers, and gene bodies. When these modifications are detected in cfDNA, they reflect the cell type of origin and dynamic biological processes. For example, H3K4me2 or H3K4me3 marks accessible active promoters, whereas H3K4me1 or H3K4me2 indicates enhancers, and H3K36me3 indicates the gene body regions of actively transcribed genes.
Studies have shown that cfDNA-associated histone modifications can be used to identify changes in liver-specific transcriptional programs in liver disease, increases in heart cells after acute myocardial infarction, and molecular heterogeneity in patients with colorectal cancer, from blood samples. These findings suggest that cfDNA-associated histone modifications have the potential to serve as powerful biomarkers for various diseases and biological processes9. cfDNA-associated histone modifications have shown promising results in distinguishing patients with colorectal cancer from healthy individuals, with an impressive AUC of 0.94. These findings demonstrate the potential of cfDNA epigenetics to provide precise diagnosis of dynamic diseases. However, further exploration of cfDNA-associated transcription factors is necessary to better understand the transcriptional regulatory relationships and support precision treatment for heterogeneous cancers. Analysis of the transcriptional regulatory relationships inferred from cfDNA might enable the identification of specific targets for therapeutic intervention and the development personalized treatment strategies for patients with cancer. Thus, continued research in cfDNA epigenetics and transcriptomics will be important, because these emerging fields have the potential to revolutionize approaches to disease diagnosis and treatment. However, the quality of the antibodies used to detect histone modifications might affect detection performance.
cfDNA transcription start site coverage indicates chromatin accessibility of gene promoter regions
cfDNA coverage at the TSS can provide insights into the nucleosome footprints and chromatin accessibility of gene promoter regions. This blood epigenetic marker accurately reflects gene activity and dynamic biological processes. In 2016, Ulz et al.13 performed whole-genome sequencing to compare sequencing coverage at TSSs in plasma samples from 104 healthy donors. A noteworthy disparity was observed in coverage between housekeeping genes and unexpressed genes, with the former exhibiting significantly lower coverage. This pattern was consistent with micrococcal nuclease data, which demonstrated a correlation between higher gene expression levels and lower coverage at TSSs. Furthermore, examination of the relationship between gene expression and TSS coverage in cancer intriguingly indicated that tumor signals could be captured from the peripheral blood by using nucleosome-depleted regions13.
Concurrently, Snyder et al.14, while investigating hematopoietic cell characteristics in blood samples, notably also identified additional fragments reflecting tumor information, which was often specific to cancer types. Through analysis of 76 RNA expression datasets, they discovered strong correlations, thus suggesting that cancer types can be accurately identified according to the non-hematopoietic contributions found in patients’ circulating cfDNA14. These findings suggest that cfDNA TSS coverage has the potential to serve as a powerful biomarker for various diseases and biological processes, thus providing a more comprehensive understanding of disease progression and aiding in the development of personalized treatment strategies. cfDNA TSS coverage also has potential clinical application in early cancer detection.
Use of cfDNA promoter fragmentation entropy to infer gene expression levels
Ongoing research continues to investigate cfDNA promoter regions. Findings have demonstrated that cfDNA PFE displays similar performance to that of cfDNA TSS coverage in correlating gene regulation activity. In fact, plasma cfDNA PFE has been found to have a strong positive correlation with gene expression levels in peripheral blood mononuclear cells (R = 0.89, P < 0.0001). Additionally, cfDNA PFE has shown promise in identifying lung cancer, with an AUC of 0.83 in the validation set, and in distinguishing LUAD from other subtypes, with an AUC of 0.9.
Furthermore, cfDNA PFE has been demonstrated to predict the response to programmed death-ligand 1 immune-checkpoint inhibition in patients with non-small cell lung cancer, with an AUC of 0.93. Patients with higher cfDNA PFE lung dynamics scores have been shown to have poorer progression-free survival (HR = 11.86, P = 0.0003).
Moreover, cfDNA PFE has the potential to dynamically monitor chemotherapy response in diffuse large B-cell lymphoma, in which the cfDNA PFE score is consistent with the circulating tumor DNA mean allele frequency and TMTV (mL), thus indicating its potential for monitoring cancer relapse events15.
New advancements in cancer detection technologies have enabled a comprehensive understanding of the genomic landscape of tumor occurrence. cfDNA-based technologies, including fragment ratio, associated histone modifications, TSS coverage, and PFE, provide enhanced insights into gene expression levels and chromatin accessibility status, thus offering valuable indications regarding the functional states of cancer cells. Moreover, they offer comprehensive information on the dynamic biological processes and heterogeneity observed during tumorigenesis. Importantly, these signatures represent a distinct departure from the focus on certain mutations and CpG islands, thus expanding understanding of cancer at a broader, more comprehensive level. Non-invasive cfDNA detection technologies in particular have great potential for exploring the dynamic biological processes of tumors, classifying cancer subtypes, evaluating response to therapy, and predicting tumor relapse.
Conflicts of interest statement
The authors have no competing interests to declare.
Author contributions
Conceived and designed the analysis: Jie Li, Xun Lan.
Wrote the paper: Jie Li.
- Received May 6, 2023.
- Accepted July 14, 2023.
- Copyright: © 2024, Cancer Biology & Medicine
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY) 4.0, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.