Abstract
Lung cancer is the most frequently diagnosed cancer and a leading cause of cancer mortality worldwide, with adenocarcinoma being the most common histological subtype. Deeper understanding of the pathobiology of non-small cell lung cancer (NSCLC) has led to the development of small molecules that target genetic mutations known to play critical roles in progression to metastatic disease and to influence response to targeted therapies. The principle goal of precision medicine is to define those patient populations most likely to respond to targeted therapies. However, the cancer genome landscape is composed of relatively few “mountains” [representing the most commonly mutated genes like KRAS, epidermal growth factor (EGFR), and anaplastic lymphoma kinase (ALK)] and a vast number of “hills” (representing low frequency but potentially actionable mutations). Low-frequency lesions that affect a druggable gene product allow a relatively small population of cancer patients for targeted therapy to be selected.
keywords
- Lung cancer
- epidermal growth factor (EGFR)
- anaplastic lymphoma kinase fusions (ALK fusions)
- tyrosine kinase inhibitors (TKIs)
- TKI resistance
Introduction
In recent years, diagnosis and treatment of patients with advanced lung cancer have undergone transformational changes. The current paradigm for prescribing novel targeted therapies is based on selecting patients according to the presence of specific oncogenic abnormalities in the tumor. The efficacy of therapy targeted at a specific oncogene is convincing evidence of “oncogene addiction”, or the concept that some cancers rely on or are “addicted to” a specific gene for their survival and proliferation1.
The first such abnormalities discovered in lung cancer were epidermal growth factor (EGFR) kinase domain mutations; tumors with these mutations were found to be sensitive to EGFR tyrosine kinase inhibitors (TKIs)2. The echinoderm microtubule-associated protein-like 4- anaplastic lymphoma kinase (EML4-ALK) fusion has emerged as the second most important driver oncogene in lung cancer and the first targetable fusion oncokinase to be identified in 4%-6% of lung adenocarcinomas3,4. Crizotinib, an oral small-molecule inhibitor of ALK and c-MET receptor kinases, is now approved for treatment of ALK positive advanced non-small cell lung cancer (NSCLC), based on the results of two pivotal studies5–7.
Among EGFR or ALK mutated NSCLCs, the percentage of complete response is negligible and therefore novel, effective, safe treatments need to be tested and developed. To this end, repurposing an existing drug for treatment of NSCLC is also a worthy goal.
EGFR NSCLC patients: second line therapy and beyond
Our group was the first to evaluate the feasibility of large-scale screening for EGFR mutations in patients with advanced NSCLC and analyze the association between the mutations and the outcome of erlotinib treatment8. Since the introduction of erlotinib and gefitinib, patients with metastatic EGFR positive lung cancer can be offered a therapeutic alternative that has proven its superiority over standard platinum-based chemotherapy2,9. In the EURTAC study, in which erlotinib was compared with platinum-doublet chemotherapy as first-line treatment for patients with EGFR-mutant NSCLC, erlotinib demonstrated a significant improvement in the overall response rate (ORR) and median progression free survival (PFS)2. On the basis of this study, in May 2013 the U.S Food and Drug Administration (FDA) approved erlotinib for use in patients with lung cancers harboring EGFR exon 19 deletions and EGFR L858R substitutions2.
Whereas reversible EGFR TKIs compete with ATP in the kinase domain of EGFR, 2nd generation EGFR TKIs, also compete for ATP binding but then covalently bind at the edge of the ATP binding cleft on Cys773 of EGFR via the Michael mechanism (addition of nucleophile to an α, β unsaturated carbonyl)10. Currently there are two lead 2nd generation EGFR TKI candidates, afatinib and dacomitinib, that are active against EGFR mutations with acquired resistance to erlotinib or gefitinib11. Afatinib exhibits superior anticancer activity in lung cancer patients harboring gefitinib/erlotinib-resistant mutant EGFR (including T790M, exon 20 insertion, and T790M/L858R double mutation)12. Based on the results of the LUX-Lung 3, the FDA has approved afatinib as a new first-line treatment for patients with metastatic EGFR-mutated NSCLC13. In the pooled analysis from LUX-Lung 3 and 6, presented this year at ASCO, afatinib prolonged survival of lung cancer patients whose tumors have common EGFR mutations by a median of three months compared with standard chemotherapy and significantly reduced the risk of death by 19%. The most pronounced reduction in risk of death was 41% in patients whose tumors had the most common exon 19 deletion EGFR mutation; for patients with the L8585R mutation there was no impact on overall survival14.
Despite the recent paradigm shift in the treatment of NSCLC patients, with a move toward biomarker-directed therapy, preclinical data has seldom been incorporated into clinical practice, since the existence of multiple resistance mechanisms complicates the selection of optimal biomarkers. Among patients progressing to first generation EGFR TKIs, 50% have tumors with a secondary T790M mutation15. The emergence of the T790M EGFR gatekeeper mutation and up-regulation of downstream signaling by MET amplification have been described as the two main mechanisms responsible for acquired resistance16. However a phase III trial enrolling only patients with MET-positive tumors was stopped in early March 2014 due to futility; there was no evidence to suggest a positive effect of addition of onartuzumab to erlotinib17.
Other mechanisms include EGFR amplifications, PI3KCA mutations or a transition from epithelial to mesenchymal differentiation. For a small percentage of resistant tumors, histological transformation occurs to small cell lung cancer (SCLC)16. We recently reported possible causes of resistance to EGFR TKIs in EGFR-mutant NSCLC patients: the high co-existence of the pretreatment somatic T790M mutation, with a clear impact on PFS, and the role of BIM mRNA expression as an independent prognostic marker18. Pretreatment T790M can be detected in more than 60% of cases18. In fact, using a PCR-PNA assay pretreatment, T790M mutations were detected in 65.26% of patients18. These results reinforce the need for 2nd and 3rd generation EGFR TKIs, while taking into account existing data that suggest use of erlotinib or gefitinib beyond progression, with added chemotherapy, radiotherapy or best supportive care may improve survival19. Although afatinib and dacomitinib have been introduced to overcome acquired resistance, they showed limited efficacy in NSCLC with T790M and were more than 100-fold less potent in NSCLC cells with EGFR T790M mutation than in NSCLC cells with EGFR activating mutation20.
CO-1686 is a novel covalent inhibitor that irreversibly and selectively targets both the initial activating EGFR mutations and the T790M secondary acquired resistance mutation21. To investigate its use as a single agent, CO-1686 is being evaluated in a phase I/II trial in EGFR-mutant NSCLC patients previously treated with first-line gefitinib or erlotinib (NCT01526928)22. In the phase I study, and based on early findings from the ongoing phase II trial, the agent yielded a 58% ORR across all dose levels in trial participants with biopsy-confirmed EGFR T790M mutations. Additionally, the compound did not cause the skin rash and diarrhea commonly associated with earlier generations of EGFR inhibitors23. AZD9291 showed high activity in preclinical studies and was well tolerated in animal models. Xenograft studies identified a breakdown metabolite of AZD9291 called AZ5104 that is about five times as potent as AZD9291 itself24. In the phase I study of AZD9291 in EGFR mutant patients resistant to standard EGFR TKIs, 50% of patients experienced tumor shrinkage and the drug worked particularly well in patients with the T790M mutation25.
A consistent proportion of EGFR mutant patients, approximately 30%, never respond to EGFR TKIs due to primary resistance. BIM mRNA expression could enrich the molecular diagnosis in patients with EGFR mutations, by identifying patients unlikely to respond and those with stable disease for whom treatment may not be effective due to early adaptive resistance18,26. It is worth mentioning that, besides BIM expression, a common BIM deletion polymorphism impairs generation of the proapoptotic isoform required for EGFR TKIs27. This polymorphism confers an inherent drug-resistant phenotype that can be circumvented with the addition of the histone deacetylase (HDAC) inhibitor vorinostat to EGFR TKIs28.
The two primary signaling pathways activated by EGFR include the mitogen-activated protein kinase (MAPK), and the phosphoinositide-3-kinase (PI3K), axes. Src tyrosine kinases and activation of the signal transducer and activator of transcription 3 (STAT3) pathways and downstream signaling have also been well documented29. EGFR phosphorylation leads to recruitment of multiple effector proteins through recognition and binding of Src-homology 2 domain-containing phosphatase 2 (SHP2) to phosphotyrosine motifs on the receptor29. SHP2 (encoded by PTPN11) is a ubiquitously expressed SH2 domain-containing protein tyrosine phosphatase (PTP). Despite its direct function in protein dephosphorylation, SHP2 plays an overall positive role in transducing signals initiated from growth factors/cytokines and extracellular matrix proteins, and initiating various downstream signaling cascades, including the PI3K and MAPK29,30. By contrast, SHP2 functions as a negative regulator of the JAK/STAT pathway31.
In 2004, Sordella and colleagues were able to demonstrate the differential EGF-induced tyrosine phosphorylation pattern seen with wild-type (WT) and mutant EGFR receptors32. For instance, Y845 is highly phosphorylated in the L858R missense mutant, but not in the WT or deletion mutant, and hence appears to be unique in distinguishing between the two types of EGFR mutations32. Y845 (pY845) phosphorylation stabilizes the activation loop, maintains the enzyme in an active state, and regulates STAT3/5 activity32. Surprisingly, the EGFR L858R mutation leads to decreased ability to activate ERK compared to WT EGFR which correlates with decreased EGFR internalization, reduced phosphorylation of SHP2, hyperactivity of STAT3 and reduced sensitivity to gefitinib33. Lazzara and colleagues found that SHP2 Y542 phosphorylation was not induced in response to EGF in the H3255 cells, which harbor the missense L858R exon 21 mutation, suggesting that SHP2 activity may be less efficiently promoted by EGFR L858R and the STAT3 pathway may be more active34. The main problem is that STAT3 activation is not abrogated by single EGFR TKI treatment with gefitinib, erlotinib, afatinib or dacomitinib20,34–36. Indeed, the second generation irreversible EGFR TKIs, afatinib or dacomitinib, not only do not abrogate, but may even induce STAT3 phosphorylation in gefitinib or erlotinib resistant cell lines such as H1975 or PC9-R20.
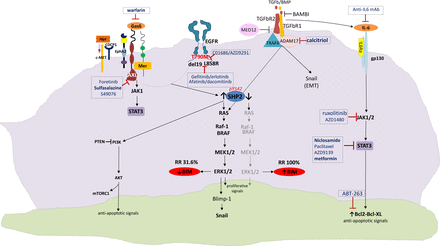
Afatinib activates interleukin-6 receptor (IL-6R)/JAK1/STAT3 signaling via autocrine IL-6 secretion in both cells. Blockade of IL-6R/JAK1 significantly increased sensitivity to afatinib through inhibition of afatinib-induced STAT3 activation. The role of the paracrine IL-6R/JAK1/STAT3 loop between stroma and cancer cells in the development of drug resistance is crucial20. Yao et al. uncovered the existence of a subpopulation of cells intrinsically resistant to erlotinib which display features suggestive of epithelial-to-mesenchymal transition (EMT) in NSCLC-derived cell lines and early-stage tumors before erlotinib treatment37. Activation of TGF-beta-mediated signaling was sufficient to induce these phenotypes. An increased TGF-beta-dependent IL-6 secretion released previously addicted lung tumor cells from their EGFR dependency. Therefore, both tumor cell-autonomous mechanisms and/or activation of the tumor microenvironment could contribute to primary and acquired erlotinib resistance, and as such, treatments based on EGFR inhibition may not be sufficient for the effective treatment of lung-cancer patients harboring mutant EGFR37. Combination of EGFR TKIs with drugs that target the IL-6R/JAK1/STAT3 pathway can be a rational synthetic lethal approach to overcome this mechanism of resistance (Figure 1).
Crosstalk signaling pathways as potential mechanisms of resistance to reversible or irreversible EGFR TKIs. EGFR, epidermal growth factor; TKIs, tyrosine kinase inhibitors.
Furthermore, tumor cells exposed to reversible or irreversible EGFR TKIs display early resistance dependent on MET-independent activation of BCL-2/BCL-XL survival signaling26. According to the study of Fan and colleagues, such cells displayed a quiescence-like state that was readily reversed after withdrawal of targeted inhibitors. BCL-2 induction and p-STAT3 (Y705) activation were found within the residual tumor cells surviving the initial antitumor response to targeted therapies26.
Although MAPK signaling is more commonly associated with receptor tyrosine kinases, it has been demonstrated that TGFBR1 also has tyrosine kinase activity. TGFBR1 directly phosphorylates tyrosine on ShcA, allowing for interaction with Grb2 and Sos. This leads to Ras activation, downstream ERK phosphorylation and BIM downregulation38. Furthermore, TGF-b can induce expression of the transcription factor Snail, which leads to repression of E-cadherin transcription. SMAD-independent activation of Snail is a result of RAS/ERK activation. Although the mechanism remains incomplete and deserves further study, in the tumor setting of aberrant TGF-b signaling, TGF-b actively represses E-cadherin and promotes EMT38.
More interesting is the fact that ADAM metallopeptidase domain 17 (ADAM17) or tumor necrosis factor alpha converting enzyme (TACE) targeting of TGFBR1 has been linked to ERK activation in nonprostate cells, resulting in shedding of the TGFBR1 ectodomain38. There is evidence that active vitamin D (1,25(OH)2D) therapy effectively inhibits ADAM17 and suppresses TGF/EGFR-pathway39. Finally, Yuan et al. recently demonstrated that a novel long noncoding RNA activated by TGF-b (lncRNA-ATB) amplifies the prometastatic effect of TGF-b via two independent mechanisms: firstly by inhibiting miR-200s, releasing activity of ZEB1 and ZEB2 and promoting EMT, and secondly by stabilizing IL11-mRNA and increasing autocrine IL-11-STAT3 signaling40,41.
BIM expression in treatment naïve cancers predicts responsiveness to EGFR TKIs, but almost 2/3 of patients have low BIM mRNA levels at baseline18. SHP2, which is downstream of both EGFR and several other tyrosine kinase receptors, is required for sustained activation of ERK and BIM downregulation42,43. Upon activation of MET by its ligand, hepatocyte growth factor (HGF) which is provided by stromal cells, EGFR signaling is dramatically altered44. Indeed, HGF enables EGFR to interact with proteins which are known to be markers of a highly metastatic phenotype like the CUB domain-containing protein-1 (CDCP1), Ephrin Type-A Receptor 2 (EphA2), but mainly AXL and Mer; these interactions cannot be affected by EGFR TKI treatment. Interestingly, STAT3 signaling can be hyperactive due to upstream pathways including not only IL-6 and JAK but also AXL, providing further opportunities for combination therapies (Figure 1).
ALK NSCLC patients: second line therapy and beyond
The novel ALK fusion is formed by a rearrangement occurring on the short arm of chromosome 2 and involves the N-terminal portion of the EML4 protein and the intracellular signaling portion of the ALK tyrosine kinase receptor45. EML4-ALK generates a transforming tyrosine kinase with as many as nine different variants identified46. ALK-dependent mitogenic signaling is largely mediated via the RAS/MAP kinase pathway as well as ALK-driven PI3K activation. The JAK/STAT3 pathway also provides essential survival signals and modulates cellular metabolism regulating the mitochondrial oxidation chain. STAT3 is activated by ALK, either directly or through JAK47. Crizotinib, an ATP-competitive aminopyridine inhibits tyrosine phosphorylation of ALK with an IC50 of 20-40 nM and response in 57% of patients with ALK-rearrangement positive lung cancer5. However, most patients develop resistance to crizotinib, typically within one to two years. Studies of ALK-rearranged lung cancers with acquired resistance to crizotinib have identified ALK fusion gene amplification and secondary ALK TK domain mutations (L1196M and G1269A) in about one third of cases48.
The success of a targeted drug is critically dependent on a specific and sensitive screening assay to detect the molecular drug target. The gold standard for detection of predictive ALK rearrangements is currently break-apart fluorescence in situ hybridization (FISH) as it is able to detect all known ALK-rearrangements and was clinically validated in crizotinib clinical trials7. However, the ALK FISH assay is fraught with technical challenges, including FISH signal instability and scoring difficulties. Although ALK FISH is clinically validated, the assay can be technically challenging and other diagnostic modalities are being explored, including immunohistochemistry (IHC) and reverse transcriptase–polymerase chain reaction (RT-PCR). Very interestingly, high frequency of ALK rearrangements in thyroid cancer from atomic bomb survivors in Japan was detected by a highly sensitive RT-PCR assay from archival paraffin blocks but was not confirmed by FISH or other methods49. Targeted resequencing has recently been proved to be a promising method for ALK gene fusion detection in NSCLC, with results correlating significantly with those from FISH, RT-PCR, and IHC50.
As with the majority of targeted agents, the tremendous excitement and enthusiasm sparked by crizotinib is tempered by the reality that a fraction of the target tumors are refractory from time of treatment initiation and most patients will eventually relapse and develop resistance after initial response. ALK kinase mutations and ALK fusion copy number gain (CNG) have been termed ALK-dominant mechanisms of resistance since tumors harboring these mechanisms presumably preserve ALK signaling despite the presence of crizotinib and are still dependent on that pathway for survival51. Data from several second-generation ALK inhibitors [ceritinib (LDK378), AP26113, (alectinib)] demonstrate response rates of 55% to 60% in crizotinib-resistant ALK positive NSCLC patients, with observed or predicted disease control rates of approximately 90%51. Ceritinib (LDK378) is an oral, small-molecule, ATP-competitive, ALK TKI52. Ceritinib achieved 56% response rate in patients previously treated with crizotinib with various resistance mutations in ALK; also in patients without detectable mutations52. On April 29th 2014, the FDA granted accelerated approval to ceritinib for treatment of patients with ALK positive, metastatic NSCLC with disease progression on or intolerant to crizotinib. The approval of ceritinib was based on the results of a multicenter, single-arm, open-label clinical trial enrolling 163 patients with metastatic ALK-positive NSCLC who had progressed on or were intolerant to crizotinib. This study is part of the ongoing clinical trial program in this patient population. Zykadia achieved an ORR of 54.6% and a median duration of response of 7.4 months based on investigator assessment53. However, while ceritinib is able to effectively suppress many crizotinib-resistant mutations, the G1202R and F1174V/C mutants are resistant to ceritinib48.
One of the questions surrounding second-generation ALK inhibitors is exactly how they will fit into the treatment landscape of ALK positive NSCLC and whether they stand a chance of outshining crizotinib at the frontline. Furthermore, encouraging early clinical results in NSCLC have demonstrated that ganetespib, a novel triazolone inhibitor of HSP90, may offer a potential strategy to target ALK inhibition by inducing substantial antitumor responses and overcoming acquired resistance in patients with ALK positive lung cancer. When combined with ganetespib, crizotinib displayed superior antitumor efficacy compared with monotherapy in H3122 NSCLC xenografts54. Recently, Richards et al. found that sensitivity of EML4-ALK variants to HSP90 inhibitors differs depending on whether the breakpoint of the fusion protein interrupts the globular TAPE (tandem atypical propeller in EMLs) domain. They determined the molecular structure of a conserved, tubulin-binding region of EML1 that reveals an unexpected protein fold55. This region is disrupted in almost 70% of EML4-ALK fusions found in patients, causing them to be sensitive to drugs that target HSP90, a cellular factor that stabilizes misfolded protein. These findings suggest that the truncation of a globular domain at the translocation breakpoint may prove generally predictive of HSP90 inhibitor sensitivity in cancers driven by fusion oncogenes55.
Other “ALK-independent” mechanisms, such as the activation of compensatory signaling pathways, may also confer resistance to targeted ALK agents. Bypass signaling, by v-Kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog (KIT) or EGFR, has also been described as a mechanism of resistance to crizotinib in ALK positive NSCLC51. EGF-induced activation of EGFR, HER2, and HER3, as well as a reduced level of ALK activation, was associated with sustained downstream signaling in the presence of ALK inhibitors, indicative of a shift in survival dependency from the ALK signaling pathway to HER family pathways in ALK-TKI–resistant cells56. The combination of an ALK inhibitor and an EGFR inhibitor induced apoptosis, further supporting the notion that the EGFR signaling pathway contributes to survival in cells resistant to ALK inhibitors56. ALK rearrangements and EGFR mutations could coexist in a small subgroup of NSCLC. Advanced pulmonary adenocarcinomas with such co-alterations could have diverse responses to EGFR-TKIs and crizotinib57. Relative phospho-ALK and phospho-EGFR levels could predict the efficacy of EGFR-TKI and crizotinib. Yang et al. recently described co-altered EGFR and ALK in a large cohort of NSCLC and found that 3.9% of EGFR mutant and 18.6% of ALK rearranged tumors have co-alterations57. Tumors harboring co-altered EGFR and ALK could have diverse responses to first-line EGFR-TKIs, which were associated with phospho-EGFR levels57. Phospho-ALK levels correlated with efficacy of subsequent crizotinib treatment. In clinical practice, we should pay attention to the specific biological behavior and corresponding management of NSCLC with dual altered EGFR and ALK genes57.
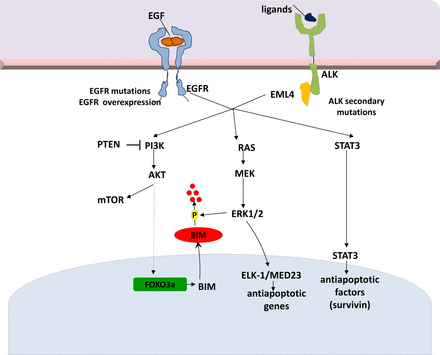
ALK and its ligand pleiotrophin are required for self-renewal and tumorigenicity of glioblastoma stem cells (GSCs)58. The pleiotrophin-ALK axis is activated by SOX2 and may be a promising target for therapy in ALK signaling-dependent tumors58. Finally, Takezawa et al. demonstrated that expression of BIM and survivin are independently regulated by ERK and STAT3 signaling pathways, respectively, and are implicated in ALK-TKI-induced apoptosis in NSCLC cells positive for EML4-ALK59. A selective inhibitor of ALK kinase activity (more potent than crizotinib), NVP-TAE68, inhibits STAT3 phosphorylation and downregulates survivin in H2228 cells, but fails to inhibit ERK phosphorylation and upregulate BIM. In a recent study, inhibition of both STAT3 and ERK pathways with the combination of TAE684 and a MEK inhibitor, AZD6244, was associated with a marked increase in the number of apoptotic cells60 (Figure 2).
EML4-ALK signaling pathways and potential mechanisms of resistance to targeted therapies. EML4-ALK, echinoderm microtubule-associated protein-like 4- anaplastic lymphoma kinase.
Conclusion
TKIs are effective anti-cancer therapies but resistance to these agents eventually develops. Several models of resistance to TKIs have been studied, including resistance to EGFR inhibitors or ALK inhibitors. A decade after the discovery of EGFR mutations, treatment is still incomplete and growing evidence indicates that targeting STAT3 or upstream or downstream components could cause induce synthetic lethal effects and achieve better outcomes. Repurposing drugs in combination with gefitinib, erlotinib, afatinib and dacomitinib is being investigated in cell cultures, subcutaneous & orthotopic xenograft models and clinical tumor samples. Clearly, the use of oncogene-addicted, highly drug-sensitive cell line models to identify escape mechanisms should be a strategy for identifying and overcoming resistance. However, it is of paramount importance that whenever possible, a biopsy is taken at time of cancer recurrence in patients who become resistant to targeted therapies so that these mechanisms can be confirmed or refuted. By identifying how a patient’s cancer becomes refractory to targeted therapies, we will be well positioned to design rational treatment strategies to re-induce remissions. Only with expanding knowledge of these escape mechanisms, availability of drugs targeting escape pathways, and examples of such molecular data informing treatment decisions resulting in good clinical outcome, can re-biopsy and molecular profiling of recurrent drug-resistant tumors be widely incorporated into clinical practice.
Acknowledgements
This work in Dr Rosell’s laboratory is partially funded by a grant from Fundació La Caixa.
Footnotes
Conflict of interest statement No potential conflicts of interest are disclosed.
- Received August 21, 2014.
- Accepted September 6, 2014.
- Copyright: © 2014, Cancer Biology & Medicine
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY) 4.0, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.