Abstract
OBJECTIVE To observe the effect of phenylacetate on the expression of RNA editing deaminase ADAR2 mRNA in glioma cells.
METHODS Primary glial cells from human brain tissue and glioma U-251 MG cells were cultured. The expression of ADAR2 mRNA was detected by a semiquantitative reverse transcription-polymerase chain reaction (RT-PCR). The levels of ADAR2 mRNA expression before and after treatment with phenylacetate were tested by RT -PCR and image analysis. The level of ADAR2 gene expression was presented as the ratio expression rate (RER) of ADAR2 gene/β-actin based on computer image analysis.
RESULTS The ADAR2 mRNA displayed mild expression in brain glial cells, and a high expression level in high-grade malignant U-251 MG glioma cells. Computer image analysis showed that the RERs of the ADAR2 gene in the U -251 MG cells before and after treatment with 4.0, and 5.0 mM phenylacetate for 8 h were 100.0, 73.5, 60.3, respectively. The expression of ADAR2 mRNA was decreased by phenylacetate in glioma U-251MG cells.
CONCLUSION Phenylacetate can decrease the expression of ADAR2 mRNA in glioma cells, suggesting that phenylacetate, as a drug, may act on the course of RNA editing in gliomas.
- glioma
- ADAR2
- RT-PCR
RNA editing takes place at a post-transcriptional level.[1] Adenosine deaminases that act on RNA (ADARs) are RNA-editing enzymes that deaminate adenosines to create inosines in double-stranded RNA (dsRNA).[2] Currently, the functions of ADAR are the focus of RNA editing research.[3,4] To date, research on ADAR2 expression in the malignant U-251MG glioma cell line has not been reported. Herein we report the mRNA expression levels of ADAR2 in the glioma cell line U-251MG and the effect of an antitumor agent, phenylacetate, on ADAR2 mRNA expression.
MATERIALS AND METHODS
Cell culture
Primary cultures of human brain astrocytes were produced. Normal human brain tissues that were used to isolate and purify astrocytes were obtained from patients who had suffered an injury of the skull and brain. The cells were routinely cultured in DMEM medium (GIB-CO, Life Technologies, Gaithersburg, MD) under 5% CO2 in humidified air at 37 °C. The primary cells were adjusted to a concentration of 106/ml. The fibroblasts were eliminated by differential adherence.
The human neuroglioma U-251MG cell line (Cell Center of Chinese Academic Medical College, Beijing, China) was used in this study. The cells were routinely cultured in the DMEM medium supplemented with supplemented with 10% fetal bovine serum/1% penicillin/streptomycin/2 mM glutamine under 5% CO2 in humidified air at 37 °C. Cell monocytes were enzymatically dissociated with 0.2%(W/V) trypsin and 0.05% (W/V) EDTA.
RNA extraction and RT-PCR
Total RNA was extracted from the culture cells using the Catrimox-14TMRNA kit(TAKARA). The ADAR2 full length primer and ADAR2 specific primer are listed in Table 1. RT-PCR was routinely performed using the RevertTMAid First Strand cDNA kit. The beta-actin primer was used as internal control (forward primer: 5’-GTT TGA GAC CTT CAA CAC CCC-3’ and antisense primer: 5’-GTG GCA TCT TGC TCG AAG CT-3’, 287 bp fragment). Each reaction system contained: 2 μl cDNA, 50 mmol/L primer, 2 |μl 2.5 mmol/L dNTPs, 8 μl 10 ? PCR buffer, 0.5 μ,l Tag plus I. Each primer was amplified using an optimized number of PCR cycles (35 cycles otherwise noted) of 94°C for 40 s, 55~65 °C for 1 min, 72 °C for 1 min, and a final extension at 72 °C for 7 min. The PCR reaction was visualized on a 1.5% agarose gel containing 5 µg/ ml of ethidium bromide.
PCR primer sequence of the ADAR2 gene
Experimental group
The cells were divided into 2 groups, including the primary human normal brain astrocytes and the neuroglioma U-251MG cells. PCRs were separately performed. The experiments were repeated 5 times. The glioma U-251MG cells were treated with 2 different concentrations of phenylacetic acid (4.0 and 5.0 mM). RT-PCR was performed to assess the expression of ADAR2 mRNA.
Digital image analysis
Semiquantitative analysis was performed using an image analyzer LUZEX-F (NIRECO, Japan). The expression level was counted using the ratio of the gray scale to the ADAR for β-actin. Data were measured 3 times.
Statistical analysis
Data were expressed as mean±SD using the Student t test. P<0.05 was considered as significant.
RESULTS
Expression of ADAR2 mRNA in glial cells of human brain tissue
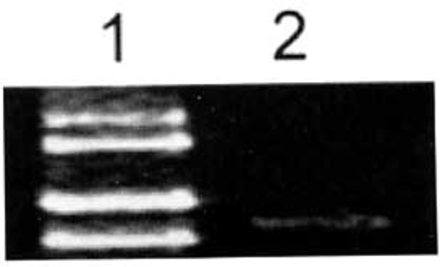
Fig. 1 shows, ADAR2 was slightly expressed in brain glial cells.
Expression of ADAR2 mRNA in glial cells of human brain tissues determined by RT-PCR. Expression of ADAR2 mRNA in glial cells of human brain was low. 1: Marker (takala F-17 λ-EcoT14 I digest); 2: Glial cells of human brain tissue.
Expression of ADAR2 mRNA in the glioma U-251MG cell line
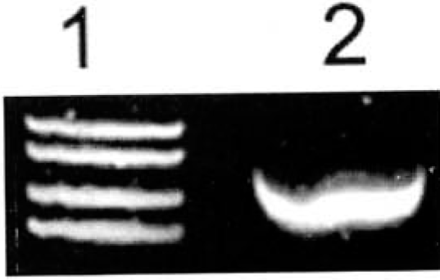
The expression of ADAR2 was strong in the highly malignant U-251MG glioblastoma cell line (Fig.2).
Expression of ADAR2 mRNA in U-251MG glioma cells determined by RT-PCR. Expression of ADAR2 mRNA was very strong in these cells. 1: Marker (takala F-17 λX.-EcoTl4 l dgest);2:Glioma U-251 MG cells.
Expression of ADAR2 mRNA in human U-251MG glioblastoma cells after treatment with phenylacetate
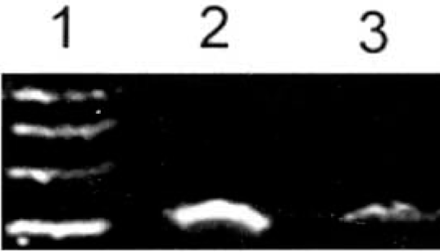
ADAR2 mRNA showed strong expression in malignant human U-251MG glioblastoma cells 8 h after treatment with 4.0 mM phenylacetate. In contrast, its expression was weak 8 h after treatment with 5.0 mM phenylacetate (Fig.3).
Expression of ADAR2 mRNA in U-251 glioma cells determined by RT-PCR before and after treatment with phenylacetate. The expression of ADAR2 mRNA was decreased by 5.0 mM phenylacetate in glioma U-251 cells. 1: Marker (takala F-17 λ-EcoT14 I digest); 2: Untreated glioma U-251 cells; 3: Glioma U-251 cells after treatment by 5.0 mM phenylacetate; 4: Glioma U-251 cells after treatment with 4.0 mM phenylacetate.
Computer image analysis showed that the ratio expression rate (RER) of the ADAR2 gene in U-251 MG glioblastoma cells before and 8 h after treatment with 4.0, and 5.0 mM phenylacetate were 100.0, 73.5 and 60.3, respectively. The expression level of ADAR2 mRNA was significantly decreased by 5.0 mM phenylacetate in the U-251 cells. This indicates that the expression of ADAR2 mRNA was significantly down-regulated by 5.0 mM phenylacetate in these cells (P<0.01).
DISCUSSION
Glioblastoma multiforme (GBM), which is the. most common malignant form of glioma, causes death in most patients within 1 year after diagnosis.[5] The U-251MG cell line was established from a human glioblastoma.[6] With regard to RNA-editing in human glioma, Mass et al.[7] in 2001, had first reported that in human malignant brain glioblastoma tissues, there was an underediting on the Q/R and R/G sites.
In mammals, large numbers of genes undergo a type of RNA editing that is characterized by site-selective adenosine-to-inosine (A-to-I) modification.[8]The A-to-I editing substrates are brain-specific transcripts coding for the glutamate receptor (GluR) channels and G-protein-coupled serotonin receptors. In the GluR subunit B(GluR-B), two editing positions, Q/R-site and R/ G-site, have been found. The GluR-B subunit mRNA was completely edited in human brain.[9,10] At the Q/R-site, RNA editing almost quantitatively (>99.9% ) changes the glutamine (Q) codon CAG to a CIG, specifying arginine (R). It is believed that the Q/R site is a naturally RNA editing site in human brain. RNA editing at the Q/R sites is “natural RNA editing”.
RNA editing is regulated by adenosine deaminases acting on RNA (ADAR).[11] ADAR-mediated RNA editing is essential for normal life and the development of both invertebrates and vertebrates.[12,13] In the ADAR family, ADAR1 knockout mice die as embryos and ADAR2 null mice are bom at full term but die prematurely. [14] ADAR2 is known as the adenosine deaminase that is primarily responsible for the Q/R site, while ADAR1 edits more promiscuously at several other adenosines in the double-stranded stem, [15] ADAR3 is capable of binding not only to doublestranded RNA (dsRNA) but also to single-stranded RNA (ssRNA).[16]
Mass et al.[7] thought that underediting on glioblastoma might be due to decreased expression of ADAR2 or down-regulation of ADAR2 enzymatic activity. However, after judging by quantitative expression analysis using real-time PCR, they found that the expression of ADAR2 mRNA was not significantly altered (not decreased) in the glioblastoma tissues.
On the other hand, in 2004, Levanon et al.[8] confirmed that abundant RNA editing occurs in neuroblastomas tissues, and the sites of these RNA editing were previously unknown. They showed that this abundant RNA editing was also regulated by ADARs. We thought that Levanon’s results suggested that ADARs or one kind of ADAR might be highly expressed in neuroblastoma. We selected the ADAR2 deaminase that is primarily responsible for the Q/R site as a target for our studies in U-251 MG cells.
In our research, we found that the level of ADAR2 mRNA expression was very low in brain glial cells and high in the U-251MG cells. Our results suggest that the high level expression of ADAR2 mRNA might be associated with genesis of malignant gliomas. We believe that there is increased RNA editing in gliomas, but this does not occur at the natural RNA editing site (Q/R site). The increased RNA editing in gliomas might be abnormal RNA editing. Our results suggest that not only the decreased of natural RNA editing, but also increased abnormal RNA editing might be involved in glioma origin.
The antitumor agent, phenylacetate (PA), can suppress the growth of glioma,[17, 18] and can also affect the course of translation in glioma cells.[19] We found that the expression level of ADAR2 mRNA reduced in the U-25 1MG glioma cells treated by phenylacetate. This suggests that phenylacetate can affect the course of abnormal RNA editing in high-grade malignant glioma cells.
Footnotes
This study was supported by the National Nature Science Foundation of China (No. 30271334).
- Received November 25, 2005.
- Accepted December 6, 2005.
- Copyright © 2006 by Tianjin Medical University Cancer Institute & Hospital and Springer