Abstract
OBJECTIVE To identify progression-related genes that lead to astrocytoma progression from a low to a high grade by analyzing the gene expression profiles from different tumor grades, and all-trans retinoic acid-treated astrocytoma cell lines.
METHODS In this study, all-trans retinoic acid (ATRA) was used to induce differentiation of SHG-44 cells. Then, by using a cDNA microarray, we compared gene expression profiling in different grades of astrocytoma cell lines (CHG-5, WHO Grade II and SHG-44, WHO Grade IV) and in ATRA-induced differentiation in SHG-44 cells. A panel of overlapped genes that might be related to tumor progression was identified, and the cell line individual variation was avoided as well.
RESULTS In the 31 overlapped genes, the stable over-expression of MDM2 and UGB as well as the repression of SOD2, G3BP, and CSTB in parental SHG-44 cells was observed and their possible roles in tumor progression were discussed. Moreover, validation of some differentially expressed genes was confirmed by Northern blots.
CONCLUSION The overlapped genes reported in this study might relate to progression of astrocytoma. Further study of progression-related genes may be helpful to explore the genetic pathways that are involved in astrocytoma progress from a low to a high grade.
keywords
Introduction
Astrocytoma, the most common type of brain tumor, is classified into 4 grades based on the level of tumor cell differentiation, proliferation potential, etc. Pilocytic (WHO Grade I) and low-grade diffuse astrocytomas (WHO Grade II) are well-differentiated tumors that typically develop in younger patients. They grow relatively slowly but have the potential to progress to anaplastic astrocytoma (WHO Grade III) and may also transform into glioblastoma multiforme (GBM, WHO Grade IV) [1]. Astrocytomas are difficult to resect thoroughly because of their invasive properties, and the brain structures critical to proper cognitive, behavioral, and motor functions are likely to be damaged during an operation. Targeted therapeutic approaches based on the knowledge of their biological progression could improve patient prognosis.
Although several genetic aberrations and gene expression changes have been shown to exist in different grades of astrocytomas[2-4], gene-expression profiling and genetic pathways associated with progression and transformation are far from being elucidated. In this study, we first compared the expression of progression-related genes between 2 different cell lines grades, i.e., CHG-5 (WHO Grade III) and SHG-44 (WHO Grade IV). In addition, all-trans retinoic acid (ATRA) was used to induce differentiation of SHG-44 glioblastoma cells. It is well known that retinoids exhibit a broad spectrum of biological activities, and participate in the onset of differentiation and the inhibition of growth in a wide variety of cancer cells[5-7]. However, data related to the effects of retinoids on astrocytomas are relatively sparse[8]. By comparing the gene expression profiling between 2 different grades of astrocytoma cells and ATRA-induced SHG-44 glioblastoma cells, a panel of overlapping genes was obtained, and the cell line individual variation was avoided as well. The possible roles of these genes and related pathways in tumor progression are discussed.
Materials and Methods
Cell culture and reagents
Two human astrocytoma cell lines, SHG-44 and CHG-5, were used in this study. The SHG-44 cell line (WHO Grade IV), which was established by Du[9], is a stable and widely used glioblastoma cell line in China[10-12]. The CHG-5 (WHO Grade II), which was set up and published by Chen et al.[12], is a glioma cell line derived from a diffuse astrocytoma. The media for SHG-44 and CHG-5 cells were Dulbecco’s modified Eagle’s and RPMI 1640 medium respectively, according to the author’s original reports. Both media contained 10% fetal bovine serum (FBS), 3% L-glutamine, 100 IU/ml penicillin, and 100 μg/ml streptomycin. The cells were cultured at 37°C in a humidified 5% CO2 atmosphere. The media for all the cultures were changed 2-3 times weekly and 24 h before collection. All the media and reagents for the cell cultures were purchased from Gibco (Gibco-BRL, New York).
Cell proliferation assay
Cells were plated in 6-well plates in triplicate at 1 × 105/ml (1.0 ml/well) and the media changed 2~3 times per week. The cells were collected daily by trypsinization (trypsin-EDTA, 1X; Gibco-BRL) untill day 7, and counted using a hemacytometer. Cell viability was monitored in parallel cultures.
Immunocytochemical staining
Cells cultured on cover slips in 6-well plates were fixed with ethanol/acetic acid (95:5) for 15 min at room temperature, washed with 1% normal calf serum (CS) in PBS for 15 min and permeabilized with 0.3% Triton X-100/PBS for 15 min. Then the cells were incubated with GFAP (monoclonal; Santa Cruz biotechnology, Santa Cruz, CA) overnight at 4°C and then washed with 0.1% Triton X-100/PBS, incubated with biotinylated anti-mouse antibody (1:100) at 37°C for 1 h, followed by incubation with fluorescein isothiocyanate-labeled streptavidin conjugate (1:100) for 1 h, washed with 0.1% Triton X-100/PBS 3 times, and mounted and analyzed with a confocal fluorescence microscope (Zeiss Axiovert 100M, Germany).
Treatment of SHG-44 cells with ATRA
According to several recent reports on the effects of ATRA on glioma cells and astrocytoma tissue[5,8,13], we adopted a widely used ATRA concentration of 10 μmol/L[8,13,14] and treated the SHG-44 cells for 3 days. Growth inhibitory effects of ATRA on the SHG-44 cells were determined by a cell-counting assay. ATRA (Sigma, St. Louis, MO) was dissolved in DMSO at 10 mmol/L under liquid nitrogen for storage. When used, the ATRA was dissolved in the above described culture media and diluted to a concentration of 10 μmol/L for cell culture.
Cell cycle analysis using flow cytometry
Flow cytometry (FCM) was used to examine the effect of ATRA on the cell cycle in ATRA-treated and parental SHG-44 cells. Briefly, the SHG-44 cells were harvested by trypsinization and adjusted to a density of about 5 × 105 cells/ml, fixed in 75% ethanol/PBS supplemented with 1% FBS, washed with PBS 3 times, stained with 20 μg/ml of propidium iodide (PI; Sigma, St. Louis, MO), treated with 10 μg/ml DNase-free RNase (Boehringer Mannheim, Germany) for 1 h, and analyzed using a FACScan flow cytometer and the CellQuest software (Becton Dickinson, San Jose, CA).
Total RNA extraction
CHG-5 cells, ATRA-treated SHG-44 cells (10 μmol/L, 3 days) and parental SHG-44 cells were prepared as described. Total RNA of the cell lines was extracted using the Trizol Reagents (Gibco-BRL) according to the manufacturer’s instruction. Briefly, about 1×107 cells were collected in 4 ml of denaturing solution. After 2 phenol/chloroform extractions, the RNA was precipitated with isopropanol, washed with 75% ethanol and air-dried. The extracted total RNA was re-suspended in RNAsecureTM solution (Ambion, Austin, Texas) and stored at –20°C. The concentration of total RNA was measured with Biophotometer (Eppendorf, Hamburg, Germany), and 200 ng RNA of each sample was used to run a 1% denatured agarose gel to verify the quality.
cDNA microarrays and data analysis
The 9,182 human genes were amplified with the polymerase chain reaction (PCR) from a Human UniGEM™ V2.0 cloned library (Incyte, Fremont, CA). The purified cDNA probes were re-suspended in water, mixed with 30% glycerol in 2 × TE buffer in 1:1, and spotted by SPBIO (MiraBio, Alameda, CA). The arrays were posttreated by baking, UV crosslinkage, chemical blocking and denaturing, to attach the DNA fragments to the glass surface.
The cDNA products of CHG-5 and ATRA-treated SHG-44 cells and parental SHG-44 cells were obtained by reverse transcription in Superscript II reverse transcriptase (Gibco-BRL ) from 150 μg of the total RNA of each sample, labeled with the fluorescent dyes Cy5 (CHG-5 & ATRA-treated SHG-44) and Cy3 (parental SHG-44), respectively, and then hybridization was carried out at 65°C overnight. The microarray imaging was scanned by GenePix 4000B and analyzed by GenePix Pro 3.0 (Axon, UnionCity, CA).
All data sets were normalized by a normalization factor from the average of every 300 spots of the total fluorescence ratio. When a gene appeared at least 2 times in the triplicate microarray hybridizations with normalized ratio of background subtracted Cy5 intensity versus background subtracted Cy3 over 2.0, or less than 0.5 (also the third value was not a contradictory trend), it was regarded as a differentially expressed gene. The normalized ratio of background subtracted Cy5 intensity over background subtracted Cy3 greater than 2.0 or less than 0.5 was considered as up-regulated and down-regulated gene expression, respectively.
Northern blots
To confirm the results of genes detected by the cDNA microarray, a few of differentially expressed cDNA clones were randomly selected from our microarray results, and were amplified by PCR individually to serve as probes for Northern hybridization. Using 1% agarose gel electrophoresis, 10 μg of the total RNA isolated from the cultured cells was transferred onto a nylon membrane (Hybond-N+; Amersham, Buckinghamshire, England). The membrane was UV-cross-linked and hybridized with 32P-labeled probes. The relative amount of mRNA was detected by autoradiography. The expression of the β-actin gene was used as an internal control.
Statistical analysis
Data were expressed as means ± SD and analyzed by the t test using SPSS 12.0 software. P < 0.05 was considered statistically significant. Each experiment was repeated 3 times with 3 replicate samples.
Results
Comparison between CHG-5 and SHG-44 cell lines
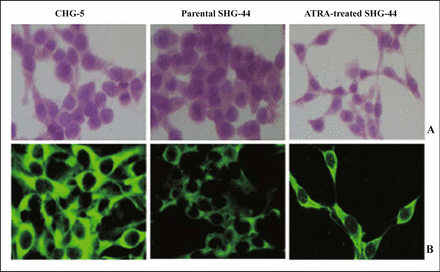
CHG-5 and SHG-44 cell lines were used for malignant grade comparison at the gene expression level. The morphology of the cell lines was not significantly different under light microscopy (Fig.1A). However, in the CHG-5 cells, the average fluorescence intensity of the glial fibrillary acidic protein (GFAP) was (45.7 ± 6.1), which is higher than that of SHG-44 cells (21.8 ± 3.1) (P < 0.05) (Fig.1B). As an often-used cell specific differentiation marker for astrocytes and astrocytomas, it is well accepted that GFAP is positively related to the level of cell differentiation[15]. The stronger GFAP expression in CHG-5 cells compared to SHG-44 cells demonstrated that CHG-5 cells are more differentiated than SHG-44 cells. A similar result has been reported by Chen et al.[12] In addition, the growth rate study demonstrated that SHG-44 cells had a higher proliferative rate than that of CHG-5 cells (Fig.2).
A, Cell morphology under light microscopy. B, Stronger cytoplasmic immunohistochemical staining of glial fibrillary acidic protein (GFAP) in CHG-5 (Grade II) and ATRA-treated SHG-44 cells compared to parental SHG-44 (Grade IV) cells under confocal microscopy.
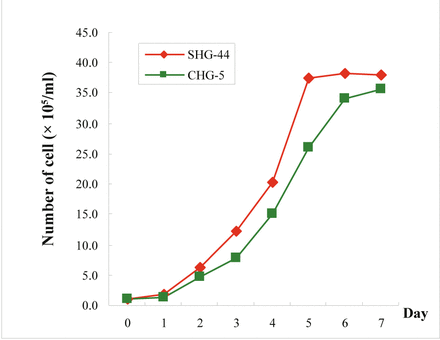
Growth curve of CHG-5 and SHG-44 cells.
Impact of ATRA on the SHG-44 cell line
Parental SHG-44 cells are ellipsoid in shape, have a small cytoplasm, big nucleus, large amount of nucleoplasm, and few short prominences under light microscopy. After treatment with ATRA for 3 days, however, the SHG-44 cells were short shuttle in shape, with a small nucleus, and decreased proportion of nucleoplasm, There was an increased formation of thin cytoplasmic processes (Fig.1A); the average fluorescence intensity of glial fibrillary acidic protein (GFAP) was 32.8 ± 4.7, i.e. higher than parental SHG-44 cells 21.8 ± 3.1 (P < 0.05) (Fig.1B); the cell number was (8.1 ± 0.2) × 105/ml, which was less than that of parental SHG-44 (12.3 ± 0.4) × 105/ml (P < 0.05). In addition, flow cytometry analysis demonstrated that in ATRA-treated SHG-44 cells, the percentage of cells in G0/G1, S, and the G2/M transition was (76.9 ± 6.4)%, (13.8 ± 1.7)%, and (9.3 ±0.9)%, respectively; whereas in the parental SHG-44 cells, the percentage of cells in G0/G1, S, and the G2/M transition was (61.9 ± 5.1)%, (25.3 ± 1.9)%, and (12.8 ±1.3)%, respectively. The results showed an evident reduction of cell growth and an increase of 15% in the G0/G1 cell population in ATRA-induced SHG-44 cells on day 3 (P < 0.05). Together, the data suggested that, apart from the differentiative-promoting effect, ATRA also inhibits cell proliferation and induces cell growth arrest.
cDNA microarray and Northern Blot
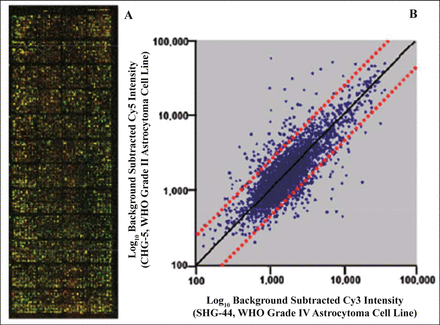
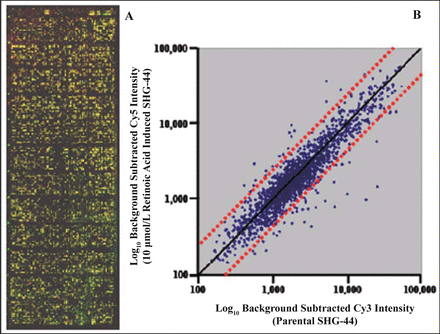
Using the 2 different grades of astrocytoma cell lines and a cDNA microarray, we first investigated 9,182 differentially expressed genes between CHG-5 and SHG-44 cells. Fig.3 shows an overlapped hybridization image of Cy5 (CHG-5, WHO Grade II) and Cy3 (SHG-44, WHO Grade IV) as well as a scatter plot of the normalized log10 intensity of the background subtracted Cy5 versus Cy3. To demonstrate the reliability of the cDNA microarray, the expression of a few genes was confirmed by Northern blots. Our results showed the same trend of the down-regulated ribosomal protein L13 (RPL13) as well as the up-regulated superoxide dismutase 2 (SOD2) determined by cDNA microarray (Fig.4). In addition, the gene expression of mammaglobin 2 (MGB2) was shown to have no significant change in both a cDNA microarray analysis and Northern blots. After treatment of SHG-44 cells with 10 μmol/L ATRA for 3 days, the ATRA-regulated genes were identified by using the same cDNA microarray. The overlapped hybridization image and normalized scatter plot of the background subtracted Cy5 intensity (ATRA-treated SHG-44 cells) versus Cy3 intensity (parental SHG-44 cells) are shown in Fig.5.
A, cDNA microarray (containing 9,182 known genes) hybridization presented the gene expression profiling of fluorescence labeled cDNA by reverse transcription from total RNA of cell lines CHG-5 and SHG-44. B, Each single spot in the scatter plot represents an expressed individual gene (B). The solid line represents the identical gene expression level in both cell lines, and the dotted lines indicate the range of 2-fold up-regulation or down-regulation.
Northern blot and microarray spot image of superoxide dismutase 2, ribosomal protein L13 and mammaglobin 2 in both CHG-5 and SHG-44 cells. The expression of actin is used for intensity normalization in the Northern blot.
A, The overlapped Cy5 and Cy3 high-density microarray image shows the expression of 9,182 genes in 10 μmol/L ATRA-induced SHG-44 cells for 3 days (Cy5 labeled) versus parental SHG-44 cells (Cy3 labeled). B, The genes located outside of the dot lines are significant differentially expressed as least 2-fold.
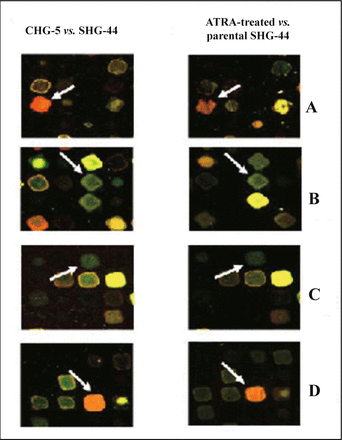
By comparing both cDNA microarray results of CHG-5 versus SHG-44 cells as well as ATRA-treated SHG-44 versus parental SHG-44 cells, 20 up-regulated and 11 down-regulated genes showed a similar expression trend. These 31 overlapped genes were shown in Table 1. Protein degradation genes such as proteasomes (PSMC1 and PSMB4) were over-expressed in CHG-5 and ATRA-induced SHG-44 cells, while the distinctive biochemical marker, intermediate filament vimentin (VIM), was strongly down-regulated. Meanwhile, the stable over-expression of MDM2, UGB as well as the repression of SOD2, G3BP, and CSTB in parental SHG-44 cells was observed. These results are the first report of the down-regulated uteroglobin gene related to malignant transformation of astrocytoma. The individual cDNA microarray spot image comparing of several genes, such as SOD2, vimentin, uteroglobin and protea-some C1 in both cell lines and ATRA-treated glioblastoma cells experiment was shown in Fig.6.
Overlapped genes comparing CHG-5 and ATRA-treated SHG-44 cells vs. parental SHG-44 cells.
A, DNA microarray images of superoxide dismutase 2; B, vimentin; C, uteroglobin and D, proteasome C1 in CHG-5 vs. SHG-44 and 10 μmol/L ATRA-treated SHG-44 cells (3 days) versus parental SHG-44 cells in a hybridization experiment.
Discussion
The Human Genome Project (HGP) was stimulated with the development of high-throughput technology for genomic study. No doubt in the near future cDNA microarrays will be one of the advanced high-throughput technologies for gene expression profiling, and it previously has been applied for defining differentially expressed genes in the brain[16], and we have used a cDNA microarray to investigate the genes that lead to progression of astrocytoma from a low to a high grade. Astrocytoma is the most common tumor of the central nervous system, and its invasive properties often make it difficult to be completely resected. Therefore, the therapeutic modalities aimed at preventing its biological progression are urgently needed.
In this study, we used 9,182 known human genes and ESTs cDNA microarray to screen genes potentially important in progression of astrocytoma from a low to a high grade. First, we compared the gene expression profiling of 2 different astrocytoma cell line grades, namely CHG-5 (Grade II, WHO) originating from a low-grade diffuse astrocytoma, and SHG-44 (Grade IV, WHO) from a glioblastoma multiforme (Fig.3). Second, in order to target the progression-related genes and get rid of the genetic variations in the cell lines, the glioblastoma cell line SHG-44 was treated for 3 days with 10 μmol/L ATRA, an antitumor and differentiation-inducing agent. The results demonstrated that ATRA promotes cell differentiation, inhibits cell proliferation and induces cell growth arrest. Meanwhile, the ATRA inductive-differentiated genes were identified using the same cDNA microarray (Fig.5). We believe that the ATRA-treated SHG-44 cells were differentiated and were changed to a lower grade status with less malignancy (Fig.1). Although there is no direct evidence to prove that the pathway of malignant progression is exactly opposite the differentiation mechanism, the overlapped genes in our cDNA microarray hybridizations were observed (Table 1), and some of them are probably important and valuable for further investigation in astrocytoma progression. To confirm the results of genes detected by the cDNA microarray, a few differentially expressed cDNA clones were randomly selected from our microarray results to perform Northern blots. The Northern analysis of a few genes such as SOD2 and RPL13 were similar to the cDNA microarray hybridization result (Fig.4). The genes with potential relation to progression of astrocytoma are reported below:
The overlapped MDM2 gene (Mdm2, transformed 3T3 cell double minute 2, a p53 binding protein), over-expression has been suggested to have a role in modulating p53 tumor suppressor activity. The MDM2 itself is a target for direct transcriptional activation by p53. By binding to the p53 protein, MDM2 oncoprotein can inhibit p53 functions as a transcription factor and induce p53 degradation[17-19]. The expression of MDM2 was lower in both CHG-5 and ATRA-treated SHG-44 cells at the RNA level. The results have shown that the ATRA treatment inhibited MDM2 expression which is similar to the expression in low-grade astrocytoma tissues.
As the intricate p53/MDM2-regulatory network is being unraveled, several other differentially expressed genes were revealed in our study, including superoxide dismutase 2 (SOD2) and cystatin B (CSTB), also were consistent with the process of the network. For example, it has been demonstrated that SOD2 is responsible for destroying radicals which are normally produced within the cells, and its expression is regulated by p53[20]. In parental SHG-44 cells, probably because of the overexpression of MDM2, its expression is repressed[21]. In addition, it was reported that over-expression of SOD2 showed to suppress tumor growth[22,23]. Cystatin B, an intracellular proteinase cathepsin inhibitor, is regarded as a potential prognostic marker in various malignant diseases. A previous study has shown that reduced expression of Cystatin B was associated with metastasis of esophageal carcinoma[24], but a recent study in colorectal cancer demonstrated an inverse correlation[25].
Another interesting overlapped expressed gene detected in our cDNA microarray experiments was uteroglobin (UGB). In our study, it was consistently up-regulated in SHG-44 cells compared with CHG-5 cells and down-regulated in ATRA-treated SHG-44 cells (Fig.6C). uteroglobin is a multifunctional, secreted protein with anti-inflammatory properties and has been reported to exert antiproliferative and antimetastatic effects in some cancer cells[26]. To our knowledge, there have been no previous reports regarding uteroglobin expression changes related with astrocytoma. Mammaglobin, another member of the uteroglobin family, has attracted wide attention because of its potential clinical marker role for the breast metastases in recent years[27,28]. As a result, we are studying uteroglobin further in different grades of astrocytoma clinical samples. It might be a potential candidate marker with probable prognostic implications.
The G3BP (Ras-GTPase-activating SH3-domain-binding protein), a protein of the Ras transduction pathway with endoribonuclease activity[29], is evolutionarily conservative and it has been reported that in comparison with 2 cell sublines (BE1, LH7) derived from a human giant cell carcinoma of the lung (PG), there was a strong expression in LH7 (non-metastatic in recipient nude mice) and a very weak expression in BE1 (100% metastatic frequency)[30]. G3BP was down-regulated in parental SHG-44 cells in our cDNA microarray. We suggest that G3BP may function to suppress metastatic potential of tumor cells.
As previously discussed, we tried to identify the genetic pathway that leads to tumor progression by comparing gene expression profiling between CHG-5 cells versus SHG-44 cells and ATRA-treated SHG-44 cells versus parental SHG-44 cells. The results may have general implications related to progression of astrocytoma for a clinical study.
Footnotes
This work was supported partly by a grant from the Sichuan Provincial Bureau of Health, China (No.050209).
- Received June 30, 2008.
- Accepted November 10, 2008.
- Copyright © 2008 by Tianjin Medical University Cancer Institute & Hospital and Springer