Abstract
OBJECTIVE This study is to investigate the prevalence of promoter CpG island methylation of O6-methylguananine-DNA methyltransferase (MGMT), mismatch repair genes (hMLH1 and hMSH2) in both tumor and serum samples of gliomas.
METHODS Methylation-specific PCR (MSP) was employed to detect promoter CpG island methylation of the MGMT, hMLH1 and hMSH2 genes in 39 samples taken from surgery and 32 samples of pretreatment serum all from the patients with gliomas.
RESULTS Promoter CpG island methylation of MGMT, hMLH1 and hMSH2 was detected and the results were 46.2%, 10.3% and 20.5%, respectively in tumor DNA of the cases with gliomas, and 40.6%, 9.4% and 18.8%, respectively in serum DNA of the cases. The methylation pattern in primary tumor and serum was found to be concordant in matched tissue and serum samples of 21 patients. In the cases with positive result of methylation for MGMT, hMLH1 and hMSH2 in tumor tissues, the results of detection for those in the paired serum sample were 77.8% (7/9), 66.7% (2/3) and 75.0 % (3/4), respectively. False positive results were not obtained in any of the patients who did not exhibit methylation. No association was found between the promoter methylation of MGMT, hMLH1, and hMSH2 genes in primary gliomas and gender, age, localization, grade of malignant or tumor stage.
CONCLUSION Promoter CpG island methylation is a frequent event in gliomagenesis. Methylation analysis appears to be a promising predictive factor of the prognosis for the glioma patients treated with alkylating drugs and a noninvasive tumor marker in serum DNA.
keywords
Introduction
Glioma is one of the caners which is most difficult to be treated. Alkylating agents are the most commonly used chemotherapeutic drugs for malignant gliomas. How to prevent and surmount the resistance of tumor cells treated by chemotherapy is a prompt problem. High level of O6-methylguananine-DNA methyltransferase (MGMT) activity in cancer cells creates a resistant phenomenon by blunting the therapeutic effect of alkylating agents and may be an important determinant of treatment failure. Epigenetic silencing of MGMT gene by promoter methylation compromises DNA repair and has been associated with longer overall survival in patients with glioblastoma who received alkylating chemotherapy with carmustine or temozolomide[1-3]. Whereas MGMT mediate resistance to DNA damage caused by alkylating agents, mismatch repair protein (hMLH1 and hMSH2) is actually required for the toxicity of several anticancer drugs. It has been established that a defect in mismatch repair is associated with resistance to many clinical common chemotherapy agents[4,5].
Promoter CpG island hypermethylation is a frequent epigenetic event in many human cancers. It is a potential pathway for tumor suppressor gene inactivation. The profile of gene hypermethylation is different for each type of cancer. Promoter methylation of MGMT, hMLH1 and hMSH2 genes in 10%~60% of gliomas has been reported, but absent in normal tissues[6-8]. Furthermore, hypermetylation of MGMT and hMSH2 seems to be an early event in gliomagenesis.
Recently, hypermethylated genes have also been detected in DNA from plasma, serum, urine, sputum, faeces by methylation-specific PCR (MSP)[9-11]. As this method can detect as little as one methylated gene per 1,000 unmethylated copies, it has sufficient sensitivity to detect low concentrations of tumor DNA in serum. Identification of the occurrence of methylation for related genes in serum might provide a potentially powerful system of biomarkers for cancer treatment[12]. Therefore, in this study, we examined the prevalence of promoter CpG island methylation of MGMT, hMLH1 and hMSH2 in tissues and serum of patients with gliomas.
Materials and Methods
Patients and tissue samples
The tumor tissues of gliomas were obtained from surgically resected specimens taken from 39 patients diagnosed and treated in No. 1 Affiliated Hospital, PLA General Hospital, Beijing, China, from March 2003 to February 2008. Pretreatment peripheral blood samples were obtained from 32 patients, including 21 paired tumor tissue and blood samples. All samples were primary tumors. The histological type and grade of tumor were classified according to WHO Central Nervous System tumor classification criteria. Fresh tissue samples were stored in liquid nitrogen and serum samples were stored in -20°C. In addition, 6 normal tissue samples from patients with cerebral trauma were as the controls. Another 30 serum samples taken from normal individuals were also used as the controls for serum DNA analysis. Informed consent was obtained from all individuals for the collection of tissue or blood.
DNA extraction
Genomic DNA was isolated from freshly frozen resected specimens of gliomas (DNA Extraction Kit, Promga). Serum DNA was extracted from 500 μl of pretreated serum using a QiAamp Blood DNA Mini kit (QiaGen) according to the manufacturer’s instructions.
Methylation-specific PCR (MSP)
The genomic and serum DNA were treated with sodium bisulfite before MSP (EZ DNA Methylation-goldTM Kit). After the treatment, the methylated cytosine remained intact while the unmethylated cytosine was completely converted into uracil which was detected as thymine following PCR.
The treated DNA was suspended in 20 μl of Tris Buffer and subjected to MSP using unmethylation-specific or methylation-specific primers. The three genes were analyzed invidividually. Promoter sequence, annealing temperature and the size of expected product are listed in Table 1.
Three genes MSP Primer sequences, annealing temperatures, and PCR product sizes.
PCR amplification was performed using Hot Start Taq DNA polymerase (Takara) according to the manufacturer’s protocols. Thermal cycling was initiated at 95°C for 5 min, followed by a series of procedures: 35 cycles of 95°C for 45 s, specific annealing (Table 1) for 45 s, extension at 72°C for 60 s; and a final extension at 72°C for 5 min. For the amplification of serum DNA, the process described above was repeated for 40 cycles. DNA from normal peripheral blood lymphocytes (PBL) was used as a control for the unmethylated promoter. Distilled water was used as negative control. A methylation-positive DNA control was made from PBL in vitro using SssI methylase (New England Biolabs) which methylated every cytosine of CpG dinucleotide in the DNA. The PCR products were fractionated by 3.0% agarose gels electrophoresis and stained with ethidium bromide.
Statistical analysis
All the statistical analyses were performed with the software SPSS 13.0 for windows. Numerical and categorical data were analyzed for statistical significance using t test and chi-square analysis, respectively. Parameters including age, histology type, stage (early vs. late), grade of malignancy (I~II vs. III~IV), and methylation status of the three genes were regarded as multivariate to be analyzed using logistic regression. A P value less than 0.05 was taken as significant.
Results
Prevalence of promoter CpG island hypermethylaiton in the tissue samples of gliomas
Methylation of the three genes was detected in the tumor samples but not in any of the normal tissue samples. The prevalence of methylation of the three genes in 39 gliomas tissue is summarized in Table 2.
Prevalence of promoter CpG island hypermethylaiton in the tissue and serum samples of gliomas patients (%).
Methylation was detectable in, at least one gene, in 64.1 % (25/39) of gliomas. Of these, 17.9% (7/39) of the cases were methylation positive in two genes, with another 5.1% (2/39) of the cases positive in all three genes. Methylation was more frequently detected in MGMT (46.2%, 18/39) than that in hMLH1 (10.3%, 4/39) and hMSH2 (20.5%, 8/39).
Table 3 illustrates the distribution of methylation of the three genes according to clinic pathological parameters of age, gender, stage, and grade. The prevalence of methylation in MGMT, hMLH1 and hMSH2 showed no significant difference between early stage and late stage (P > 0.05); or between less malignant (I~II) and high malignant (III~IV) cancer (P > 0.05). The results also indicated that the methylation of MGMT, hMLH1 and hMSH2 was not associated with age or gender of the patients. However, by both univariate and multivariate analyses, occurrence of methylation of the MGMT gene was associated with the occurrence of methylation of the hMSH2 gene (P = 0.017 and P = 0.039, respectively).
Prevalence of methylation of different genes in tissues of gliomas patients and relation to clinical data (%).
Prevalence of promoter CpG island hypermethylation in serum samples of gliomas
The prevalence of methylation of MGMT, hMLH1 and hMSH2 in serum was 40.6 % (13/32), 9.4% (3/32) and 18.8% (6/32), respectively. Methylated sequence of at least one gene was detected in 56.3 % (18/32) of serum samples. In contrast, methylated DNA was not detectable in any of the 30 normal serum samples. A higher percentage of methylation of MGMT gene in serum samples was detected compared with that in hMLH1 and hMSH2 (Table 2).
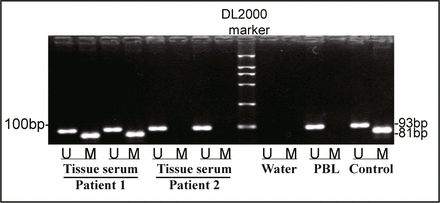
Of the 32 serum samples obtained, 21 of them had paired tumor tissue available. We found a similar pattern of methylation changes for the three genes in the serum samples to their paired tumor tissues (Fig. 1, Table 2).
Methylation-specific PCR analysis of MGMT hypermethylation. In case 1, promoter methylation was detected in both tumor and serum samples. In case 2, promoter methylation was not detected in tumor or matched serum sample. DNA from normal peripheral blood lymphocytes (PBL) was used control for the unmethylated MGMT promoter (U), enzymeatically methylated DNA from PBL served as a positive control for the methylated promoter (M), and water was used as a negative control for the PCR. A 50-bp marker ladder was loaded to estimate molecular size, as shown on the left scale; the sizes of PCR products are indicated on the right scale.
Methylation of MGMT, hMLH1 and hMSH2 was detected in these paired tissue samples with the number of 42.9% (9/21), 14.3% (3/21) and 19.0% (4/21), respectively. Of the patients with methylaiton positive in the tissue specimens, methylaiton of MGMT, hMLH1 and hMSH2 detected in serum was 77.8% (7/9), 66.7% (2/3) and 75% (3/4), respectively. Patients without methylation in glioma tissues was found to have on methylation in their serum either. The occurrence of methylation for the three genes in serum was associated with that in their paired tumor tissues (P = 0.018, Fisher’s Exact Test).
Discussion
Alkylating agents are the most commonly used chemotherapeutic drugs for malignant gliomas. DNA repair enzyme-MGMT plays an important role in the defense against alkylating agents that generate lethal DNA adducts. MGMT activity is a major mechanism of resistance to alkylating drugs, whereas DNA mismatch repair proteins (MMR) hMLH1 and hMSH2 can signal apoptosis induced by DNA damage as part of cytotoxic effects of chemical agents. Hypermethylation of the CpG island in the promoter region is an important mechanism responsible for the loss function of DNA repair proteins and impacts on the therapeutic response to alkylating agents in glioma patients. Prediction of treatment response may allow selection of those who are most likely to benefit from receiving alkylating agent as chemotherapy.
Methylaiton of MGMT, hMLH1, and hMSH2 showed in many cancers has been reported, such as esophagus, lung, colon, and cervix. However, they have not found methylated in normal tissues including buffy coat, bronchial brush, and cervix samples from healthy subjects as well as from normal tissues of cancer patients[13-15]. Tumors release their DNA into the circulation system, and the serum and plasma of cancer patients contain more free DNA than normal individuals do. One approach for cancer detection has been to detect the alterations of tumor specific DNA in the circulating DNA of patients[16,17]. Hence, in this study, we determined the prevalence of methylation of these three genes in both tissue and serum samples from the patients with gliomas.
Consistent with others, no methylation of the three genes was detected in normal tissue in the present study. In contrast, the studies in methylation of the three genes, at least one, MGMT, hMLH1, and hMSH2 was detected in about 64.1% of gliomas. Methylation of MGMT was detected in 46.2% of gliomas. The overall findings suggest that methylation of MGMT, hMLH1, and hMSH2 occurs frequently and may be associated with the gliomagenesis. The highest prevalence of methylation among all genes was also observed in the MGMT, as demonstrated in this study. We also found that methylation of hMLH2 but not hMSH1 was more common in gliomas. This indicates that methylation of particular genes may preferentially occur in particular types of cells, and promoter hypermethylation may also be a cell-type-specific event. Great precautions should be taken when selecting methylated DNA markers for cancer detection.
Promoter hypermethylaton has been reported in the serum/plasma of various cancer patients, including colorectal cancer and gastric cancer[18,19]. However, methylation in plasma/serum samples from glioblastoma has not been reported yet. In all 32 serum samples in the study, methylation of at least one gene was detected in about 56.3% of serum. Furthermore, the occurrence of methylation for the three genes in serum was associated with that in their paired tumor tissues. And no methylated sequence was detected in serum from those negative for methylation in paired tissues, which implies an absence of false-positive methylation in this study. Thus methylated genes could also be detected in serum of glioma patients. The presence of tumor-specific methylated gene sequences in peripheral blood may be developed as a noninvasive approach for cancer detection. It is possible to detect abnormality frequently using several methylated markers in majority cases with cancers. Further analysis in a much larger size of samples is warranted.
Though in our study, no significant difference was detected in methylation of MGMT or MMR between early and late stages or grades of tumor, the significant association between MGMT and MMR methylation suggests that these genes are closely related each other during gliomagenesis. The methylation status of the MGMT, hMLH1, and hMSH2 promoter may have prognostic value and, in addition, may be a clinically relevant predictor for treatment using alkylating agents as chemotherapy.
MGMT testing for diagnosis requires sufficient and optimally preserved tumor tissue. The best results with methylation-specific PCR are obtained with cryopreserved tumor specimens, therefore, deterioration of the quality of tumor DNA induced by fixation should be avoided. Other methods, such as immunohistochemistry or MGMT activity testing, may not be reliable, since MGMT expression is prone to be induced by glucocorticoids, ionizing radiation, and genotoxic agents when the MGMT promoter is not methylated[20].
Determination in the status of promoter methylation of MGMT or mismatch-repair (hMLH1, hMSH2) by methylation-specific PCR may allow the selection of patients who are most likely to benefit from alkylating agents treatment. For patients whose tumors are not methylated at the MGMT and methylated at the mismatch-repair promoter, alternative treatments with a different mechanism of action or methods of inhibiting MGMT or expressing MMR should be developed. Stratification which is generated according to the status of promoter methylation of MGMT and MMR may be considered in which alkylating agents are used. Methylation analysis appears to be a promising prognostic/predictive factor in primary glioma and a noninvasive tumor marker in serum DNA.
Footnotes
This work was supported by a grant from National Natural Science Foundation of China (No.30671995).
- Revision received December 3, 2008.
- Accepted January 19, 2009.
- Copyright © 2009 by Tianjin Medical University Cancer Institute & Hospital and Springer