Abstract
OBJECTIVE To investigate gene mutations of epidermal growth factor receptor (EGFR) and K-RAS (Kirsten rat sarcoma viral oncogene) in Chinese patients with non-small cell lung cancer (NSCLC), and study the correlation with its protein expression and its clinical significance on gefitinib.
METHODS Detect the EGFR and K-RAS gene mutations status by gene sequencing and use the method of immunohistochemistry to detect EGFR and K-RAS protein expression.
RESULTS The frequency of EGFR mutations was 33%, mainly located in exon 19 and exon 21. The frequency of K-RAS mutations was 5.5%, mainly located in codon 12. There was no case which both had EGFR and K-RAS mutations, suggesting a mutually exclusive relationship between the two. EGFR mutations are more common in adenocarcinomas (particularly those with bronchioloalveolar features), nonsmokers and females. 16% were detected EGFR positive expression and had no correlation with EGFR mutation (P > 0.05), but had significant correlation with mutation in exon 19 (P < 0.05). The frequency of K-RAS positive expression was 52.5% and had no correlation with K-RAS mutation (P > 0.05). Twelve (8 cases were protein-negative) out of 15 gefitinib-treated NSCLC patients with disease control carry EGFR mutations.
CONCLUSION EGFR protein expression has some correlation with exon 19 mutations. Combined detection of EGFR and K-RAS gene mutations can help clinicians to choose patients who may benefit from EGFR tyrosine kinase inhibitor (EGFR-TKI) and to predict the response and prognosis of gefitinib.
keywords
- Epidermal growth factor receptor (EGFR)
- Kirsten rat sarcoma viral oncogene (K-RAS)
- Non-small cell lung cancer (NSCLC)
- EGFR tyrosine kinase inhibitor (EGFR-TKI)
Introduction
In recent years, the morbidity and mortality of lung cancer has a rapidly increased and it has become the first malignant tumor in China and even the world. Despite advances in treatment, the 5-year survival rates for all stages of lung cancer have not improved. EGFR expression occurs in many malignant tumors (including NSCLC), and has a close relationship with the proliferation of tumor cells, tumor angiogenesis, adhesion and invasion, metastasis and tumor cell apoptosis and other processes. Therefore the EGFR signaling pathway has become a research hotspot of lung cancer targeted therapy in recent years.
Much research showed that EGFR-TKI (tyrosine kinase inhibitor) have great effects in NSCLC especially in lung adenocarcinoma. And more clinical studies confirmed that the efficiency of EGFR-TKI was associated with EGFR mutations[1,2]. But the relationship between the expression of EGFR protein and EGFR-TKI, EGFR gene mutations are still controversial[3-5]. K-RAS plays an important role in the downstream flow of EGFR signal transduction pathway and its mutation can lead to this pathway being continuously activated, which finally results in EGFR-TKI primary resistance. But the relationship between expression of K-RAS protein and gene mutation has been little researched and needs further investigation.
Materials and Methods
Patients and samples
Participants were 200 randomly chosen NSCLC patients who underwent surgery at Tianjin Medical University Cancer Hospital in 2009, the fresh tumor tissue was collected and frozen for preservation. Two pathologists confirmed the diagnosis of all samples which were characterized by H&E stain, and made sure that they contained more than 50% of the tumor component in order to be used in DNA extraction and mutation analysis.
DNA extraction, PCR amplification and sequencing
From each sample 25-30 mg of frozen tissue was shredded using scissors. Then E.Z.N.ATM Tissue DNA Kit (purchased by OMEGA) was used to extract DNA. Researchers then took 1 μl DNA samples from each to confirm their quality and concentration; they were then diluted to working concentration (5-10 ng/μl).
The exon 1 of K-RAS and 4 exons of EGFR (18, 19, 20 and 21) were amplified by PCR. The Forward and Reverse primers are shown on Table 1. A total of 20 μl PCR reaction system includes: 1 × HotStarTaq buffer, 2.0 mM Mg2+, 0.2 mM dNTP, 0.2 μM of each primer, 1U HotStarTaq Polymerase (Qiagen Inc.) and 10ng DNA template. PCR reaction procedures were performed using 35 cycles of 15 s at 94°C, 30 s at 56°C, 1 min at 72°C and extension for 2 min at 72°C. Sequencing reactions were electrophoresed on an ABI3700 genetic analyzer after PCR products were purified. Sequence variations were determined using Seqscape software (Applied Biosystems) with the K-RAS and EGFR reference sequence (NM_004985 and NM_005228.3, National Center for Biotechnology Information).
Specific primers of EGFR 18-21 exons and K-RAS exon 1 in PCR detection.
Immunohistochemistry
All specimens were fixed in 10% formaldehyde, embedded in paraffin, 4μm slices and dried. The immunohistochemical staining used EnVision method. EGFR staining was done in cytoplasm and membrane. EGFR results were scored as follows: 0, no staining; 1+, faint, partial membrane staining; 2+, weak, complete membrane staining in >10% of tumor cells; 3+, intense complete membrane staining in >10% of tumor cells. K-RAS staining was done in the nucleus. The percentage of positive cells were scored as follows: 0, no positive cell; 1, ≤ 25%; 2, 26%-50%; 3, 51%-75%; 4, > 75%. The intensity of staining depends on the majority of positive cells: 0, no staining; 1, weak; 2, the middle intensity; 3, intense staining. The two indicator scores were added together, and according to the total scores divided all cases into four groups: (-), 0-1; 1+, 2-3; 2+, 4-5; 3+, 6-7. Tumors with a score 2+ or 3+ were interpreted as positive for overexpression (EGFR and K-RAS).
Statistical analysis
The statistical analyses of categorical variables were performed using Pearson’s χ2 test or Fisher’s exact test, as appropriate. Two-sided P values < 0.05 were considered significant. All analyses were performed by SPSS 17.0 statistical software.
Results
Gene mutation analysis
Mutations of EGFR were detected in 66 cases (33%) out of 200 cases: 3 cases in exon 18, 37 cases in exon 19, 2 cases in exon 20 and 27 cases in exon 21. There were 3 cases (2 cases of mixed adenocarcinoma with BAC, 1 case of adenosquamous carcinoma) carrying mutations in 2 exons.
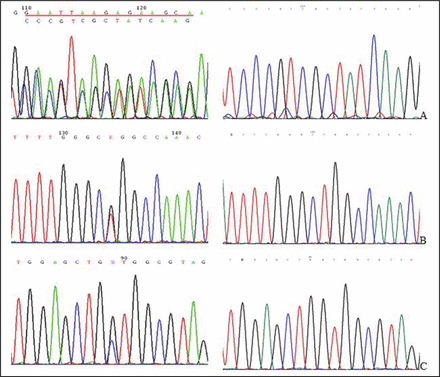
K-RAS mutations were detected in only 11 out of 200 patients (5.5%) and were all point mutations. Among all the mutations, there were 10 cases located in condon 12 and 1 case in condon 13. Sequencing analysis of gene mutations are shown in Fig. 1.
Sequencing analysis of gene mutation (the left shows mutation, the right shows wild-type). A, E746-A750 deletion mutation in exon 19; B, L858R point mutation in exon 21; C, G12A point mutation in 12 codon of K-RAS exon 1.
It was never found that both carry the EGFR and K-RAS gene mutations in 2 different samples.
Relationship between the gene mutations and clinicalpathological features, Table 2
The comparative analysis of EGFR, K-RAS gene mutations and protein expression with clinicopathological parameters (χ2 test).
In the total 200 cases of NSCLC, the rate of EGFR mutations was 33%, among these adenosquamous carcinoma reached 87.5%, adenocarcinoma 50.5%, significantly higher than squamous cell carcinoma (2.9%), large cell carcinoma (11.1%) and sarcomatoid carcinoma occupied 0% (P < 0.05). The mutation rate of BAC and adenocarcinoma with BAC was 61.3%, significantly higher than adenocarcinoma without BAC 27.8% (P < 0.05). 51.8% female patients had mutations, significantly higher than male patients 19.7% (P < 0.05). The mutation rate of people aged <65 (years) was 41.2%, significantly higher than people aged ≥ 65 (years) 17.4% (P < 0.05). 52.5% of non-smokers had mutations, significantly higher than smokers (20.0%) (P < 0.05). 32.9% of lymph node metastasis cases had mutations; there was no statistically significant difference with cases without lymph node metastasis 33.0% (P > 0.05).
Among the 11 (5.5%) patients who experienced K-RAS mutations, 9 patients were ever-smokers and 8 patients were male, 7 patients had adenocarcinoma, 2 patients had sarcomatoid carcinoma, 2 patients had squamous cell carcinoma.
Immunohistochemistry analysis
Thirty-two cases (16%) showed EGFR positive expression. Among these 25% of non-smokers were positive, and only 10.0% of smokers were positive (P < 0.05). Among the 32, 37.5% of those with adenosquamous carcinoma were postivie, 18.0% with adenocarcinoma, 10.3% with squamous cell carcinoma, 0% with large cell carcinoma and 50% of patients with sarcomatoid carcinoma (P < 0.05). The relationships with age, gender, and lymph node metastasis were not significantly correlated (P > 0.05). This is shown in Table 2.
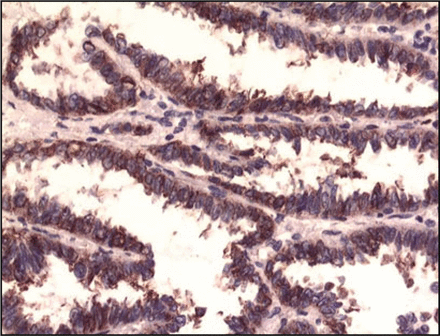
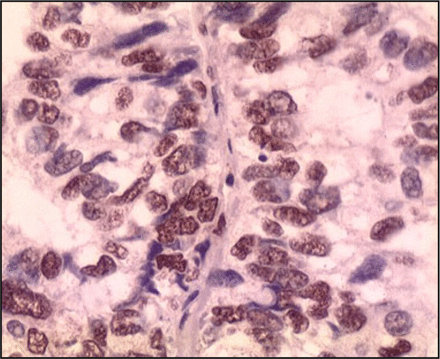
One hundred and five (52.5%) out of 200 cases showed overexpression of K-RAS protein. And the correlation with histology types, age, gender, smoking history, and lymph node metastasis were not significantly correlated (P > 0.05). The immunohistochemistry staining is shown in Fig. 2 and 3.
Expression of EGFR in non-small cell lung cancer tissues (SP × 400).
Expression of K-RAS in non-small cell lung cancer tissues (SP × 400).
The relationship between protein expression and gene mutations
The EGFR protein expression had no correlation with EGFR total mutations and exons 18, 20, 21 (P > 0.05); but had a significant correlation with exon 19 (P < 0.05). The K-RAS protein expression had no correlation with gene mutations (P > 0.05).
Gene mutations, protein expression and gefitinib sensitivity
Investigation was done into 25 patients who received gefitinib treatement randomly, it was found that 15 cases exhibited EGFR mutations. Among the patients who exhibited mutations, 12 cases (8 cases EGFR protein was negative and 3 out of 4 cases with protein overexpression had exon 19 mutation) benefited from gefitinib (80%); but only 2 out of 10 cases without EGFR mutations benefited (20%). The two groups had a statistically significant difference (P = 0.005). This is shown in Table 3.
The analysis of clinical curative effects in 25 gefitinib-treated NSCLC patients.
Discussion
More clinical studies confirmed that the efficiency of EGFR-TKI is significantly associated with EGFR mutations, and the response rate can reach 65-83% for patients with EGFR mutations, but only 10%-15% for patients with wild-type. Tamura et al.[1] studied the IIIB/IV NSCLC patients who harbored EGFR mutations and found that the overall response rate was 75%; the DCR (disease control rate) was 96%; the median PFS (progression-free survival) was 11.5 months; the median OS (overall survival) has not yet been reached, and the 1 year survival was 79%. In our research the overall response rate was 80% for patients with EGFR mutations who received gefitinib, but only 20% for patients with wild-type. All these confirm that EGFR mutations are very important for choosing EGFR-TKIs.
In our research, the rate of EGFR mutations was 33%, greatly higher than North-America (3%-25%) and South-Europe (10%-24%)[6]. The mutations mainly focused on exons 19 (56%) and 21 (41%). In exon 19, the mutations generally were single in-frame deletions and in exon 21 all mutations were L858R. The clinicopathologic characteristics that were associated significantly with EGFR mutations were sex, smoking history and pathologic type: being female, a non-smoker and having adenocarcinoma (particularly BAC and adenocarcinoma with BAC) showed a higher percentage of EGFR mutations, this was in agreeance with past reports[7]. The research showed that EGFR mutation rate of patients who < 65 years obviously higher than those who ≥ 65 (P < 0.05), it means the younger patients had higher EGFR mutation rate and more sensitivity for EGFR-TKIs. In addition, we found that the EGFR mutations rate of adenosquamous carcinoma reached 87.5%, higher than adenocarcinoma (50.5%), this possibly because the number of adenosquamous carcinomas were few (8 cases). Kang et al.[8] separated each components from adenosquamous carcinomas to investigate the status of EGFR mutation separately and concluded that : the frequency of the EGFR mutation and clinicopathologic characteristics of the EGFR mutants of adenosquamous carcinoma were similar to those of Asian patients with adenocarcinomas; identical EGFR mutations in both adenocarcinomatous and squamous cell carcinomatous components suggest the possibility of monoclonality in the histogenesis of adenosquamous carcinoma. This indicates that the treatment of EGFR-TKIs for patients with adenosquamous carcinomas can refer to adenocarcinoma.
EGFR overexpression in NSCLC has been reported always above 50%, but in our research it was 16%, similar to Dacic[9] (15.9%). This difference in values likely reflects the use of a variety of antibodies, protocols, and interpretation criteria, as well as subjectivity in scoring. The EGFR protein expression can be able or not to guide the use of EGFR-TKIs is complex and remain controversial. More studies showed that EGFR protein expression had no relationship with EGFR gene mutations[10-12]. But also have report showed that expression of EGFR were associated with an objective response of EGFR-TKIs (P = 0.03)and the overexpression group had longer survival than the negative group[4]. And before the correlation between EGFR gene mutations and EGFR-TKIs was discovered, the clinical doctors used EGFR-TKIs mainly referring to the EGFR protein expression. The EGFR protein expression had no correlation with EGFR total mutations and exons 18, 20, 21 (P > 0.05); but had significant correlation with exon 19 (P < 0.05) in our research. Among 12 NSCLC patients (EGFR mutation-type) who benefited from gefitinib, 8 cases EGFR protein expression were negative and the other 4 cases were positive, in addition, among the patients who had no response to gefitinib, there had 2 positive cases. In this situation if we depends on EGFR protein expression to choose EGFR-TKIs, there will have 8 patients who theoretically effective can not accept gefitinib treatment, but have 2 patients who theoretically having no effects will accept over-degree treatment. This will result in waste of medical resources and damage patients. Therefore only when the detection of EGFR gene mutation was confined or failed, the EGFR protein expression can use to guide the use of EGFR-TKIs at some levels.
K-RAS can result in EGFR-TKIs primary resistance and predicts a poor prognosis[13]. In our study, K-RAS mutation rate was 5.5%, obviously lower than the westerns 19%[14]. And we found that K-RAS mutations were more easy to happen in men and smokers, which is according to the before reports. No case both have EGFR and K-RAS mutation which means the two mutations are repulsive, which is agree with ever report[15]. We detected the K-RAS protein expression rate was 52.5%, and had no correlation with K-RAS gene mutation (P = 0.27).
At the present, the use of EGFR-TKIs mainly refers to: EGFR gene mutation status, EGFR gene copy number and EGFR protein expression. The front two have already confirmed by more experiments which have significant correlation with the sensitivity of EGFR-TKIs. But the correlation between EGFR protein expression and EGFR-TKIs is complex and remain controversial and needs further research.
Conflict of interest statement
No potential conflicts of interest were disclosed.
- Received March 8, 2010.
- Accepted April 10, 2010.
- Copyright © 2010 by Tianjin Medical University Cancer Institute & Hospital and Springer